MYCOBACTERIA PRODUCE PROTEINS INVOLVED IN BIOFILM FORMATION AND
GROWTH-AFFECTING PROCESSES
J
OANAK
ORABLIOVIENĖ1,2*, M
YKOLASM
AURICAS1,
ČESLOVA
A
MBROZEVIČIENĖ3,M
INDAUGASV
ALIUS4, A
LGIRDASK
AUPINIS4, S
AULIUSČAPLINSKAS5,6and P
AVELK
ORABLIOV11Department of Immunology, State Research Institute, Centre for Innovative Medicine, Vilnius, Lithuania
2Department of Epidemiological Surveillance, Centre for Communicable Diseases and AIDS, Vilnius, Lithuania
3Department of Bacteriology, National Food and Veterinary Risk Assessment Institute, Vilnius, Lithuania
4Vilnius University Life Sciences Center, Institute of Biochemistry, Proteomics Center, Vilnius, Lithuania
5Faculty of Social Policy, Mykolo Romerio University, Vilnius, Lithuania
6Centre for Communicable Diseases and AIDS, Vilnius, Lithuania
(Received: 17 July 2017; accepted: 31 May 2018)
The aim of this study was to determine the effect of mycobacterial proteins on mycobacterial biofilm formation and growth processes. We separated growth- affecting proteins (GEPs) from wild type ofMycobacterium bovisand ATCC strain ofMycobacterium avium subsp. avium. Our results showed that these mycobacteria- secreted GEPs are involved in biofilm formation, growth stimulatory, and inhibitory processes. Ourfindings suggest that GEP stimulatedM. avium subsp. aviumgrowthin vitro. Stimulation process was observed in mycobacteria affected with GEP extracted fromM. avium subsp. avium. We found that both GEPs inhibited the growth of the M. bovis. Optical density measurement and visual analysis confirm that GEP plays an important role in biofilm formation process. Most ofM. bovisGEP are associated with the type VII secretion and general secretion pathways. Our results contribute to a better understanding of the mechanisms underlying mycobacterial biofilm formation and growth-affecting processes and better characterization of mycobacterial proteins and their functions. It is noteworthy that thisfinding represents thefirst demonstration of GEP-mediated growth effects on a solid and liquid medium.
Keywords: mycobacterial proteins, mycobacterial biofilms, mycobacterial growth processes
*Corresponding author; E-mail:tamkeviciute.joana@gmail.com
Introduction
The ability of bacteria to communicate and behave as a group for social interactions like a multicellular organism has provided signi
ficant bene
fits to bacteria in host colonization, formation of bio
films, defense against competitors, and adaptation to changing environments [1]. Many bacteria have been found to regulate diverse physiological processes and group activities through a mechanism called quorum sensing (QS) [2].
With the emergence of drug resistance, treating mycobacterial infections is becoming increasingly dif
ficult and hence, looking for newer drug targets, especially those involving QS, is an essential component of mycobacterial research. However, the Gram-positive mycobacteria remain a mystery with no clear evidence known about their QS mechanism [3]. Bioinformatics analysis has revealed the presence of LuxR homologs in Mycobacterium tuberculosis, but the experimental supports are lacking [4,
5]. Some of these homologs are ubiquitousacross the multiple mycobacterial species and are involved in mycobacterial bio
film formation or persistence, suggesting a possible existence of similar QS mechanisms. Given the fact that bio
film formation is mostly linked with QS regulation [3], the existence of QS in mycobacteria cannot be ruled out. However, this hypothesis needs experimental validation [6].
M. tuberculosis typically forms pellicles at the liquid
–air interface in growth media. In recent times, pellicles have been equated to bio
films, because they are held together by extracellular polymeric substance (EPS) produced by the bacterium [7]. M. tuberculosis forms bio
films harboring antibiotic-tolerant bacilli in vitro, but the factors that induce bio
film formation and the nature of the extracellular material (ECM) that holds the cells together are poorly understood, polysaccharides, proteins, DNA, and lipids are important components of the ECM [8,
9]. However, the composition of the mycobacteria biofilm EPS and the mechanisms governing its formation remain poorly understood [9]. It is known that proteinaceous components include cell surface adhesins, protein subunits of
flagella, and pili, secreted extracellular proteins, and proteins of outer membrane vesicles [10]. Better characterization of the proteinaceous components structure, functions, and regulatory circuits controlling bio
film matrix production will provide better understanding of mycobacteria physiological processes, such as host colonization, defense against competitors, and adaptation to changing environments (e.g., antibiotic resistance). Understanding these mechanisms and their controlled social activities may open a new avenue for controlling myco- bacterial infections [1,
6,10]. In this study, we determine the effect of mycobac-terial proteins on mycobacteria bio
film formation and growth processes. We characterize these proteins by their gene name, status of existence, molecular
406 KORABLIOVIENĖET AL.
Acta Microbiologica et Immunologica Hungarica 65, 2018
weight, location, function, superfamily, and secretion pathway. Biggest part of these proteins were associated with the type VII secretion (T7S) pathway.
Materials and Methods
Bacterial strains and GEP preparation
Wild type of Mycobacterium bovis and ATCC strains of Mycobacterium avium subsp. avium (ATCC 15769) and Mycobacterium terrae (ATCC 15755) were used throughout these studies. GEPs were extracted from M. bovis and M. avium subsp. avium and tested in vitro: MA GEP
–GEP extracted from M. avium subsp. avium; MB GEP
–GEP extracted from M. bovis. Cultures were centrifuged (at 4 °C for 45 min at 4,000 rcf) after 8 and 16 weeks of incubation and passed the
filtrate through a low protein-binding 0.2-
μm
filter (Dismic-13 CP cellulose acetate
filters, Advantec, Tokyo, Japan). Concentration of proteins (CP) was quanti
fied by Bradford assay.
Growth of bacteria
Bacterial cultures (10
5CFU/ml) were transferred on Lowenstein
–Jensen medium with pyruvic acid (Becton, Dickinson and Company,
http://www.bd.com/europe/regulatory/Assets/IFU/Difco_BBL/244420.pdf). Cultures were affected by
Blank Paper Disks (6 mm diameter, Becton, Dickinson and Company) impreg- nated with GEP and incubated at 37 °C for 8 weeks. At the end of incubation, the number of bacteria colonies was calculated. In total, 100 samples were prepared.
Bio
film formation
To evaluate the effect of GEP on bio
film formation, bacterial cultures were raised in 15-ml screw-capped bottles with 2 ml of culture, 5 ml of media, and 0.5 ml of GEP. At the end of third week of incubation, the caps of bottles were loosened to allow further growth of Mycobacterium at the interface. Cultures were incubated at 37 °C for 6 weeks.
Congo red assay and cellulose optical density (OD) measurement
About 2% of Congo red was added to both the control and test samples and
continued shaking at 37 °C for 2 h. After 2 h, control and mycobacterium bio
film
cells were centrifuged at 5,000 g for 5 min, washed three times with PBS, and then were analyzed visually for Congo red binding. OD measurement was performed at 500 nm.
Filter-aided protein sample preparation (FASP)
Proteins were concentrated on Amicon Ultra-0.5 mL 30 kDa centrifugal
filter. Trypsin digestion was performed according to a modi
fied FASP protocol as described by Wisniewski et al. [11]. Brie
fly, proteins were washed with buffer containing 8 M urea. The proteins were alkylated using iodoacetamide. Buffer was exchanged by washing twice with 50 mM NH
4HCO
3, and proteins were digested overnight with TPCK Trypsin 20233 (Thermo Scienti
fic, USA). Then, peptides were recovered by centrifugation and washed with 20% CH
3CN. Afterward, samples were lyophilized, redissolved in 0.1% formic acid, and analyzed by mass spectrometry (MS).
Liquid chromatography (LC) and MS
The liquid chromatography (LC) separation of trypsin-cleaved peptides was performed with nanoAcquity UPLC system (Waters Corporation, UK). Peptides were loaded on a reversed-phase trap column PST C18 (Waters Corporation) at a
flow rate of 15 ml/min using loading buffer of 0.1% formic acid and subsequently separated on HSS-T3 250 mm analytical column (Waters Corporation) in 30-min linear gradient (A: 0.1% formic acid, B: 100% CH
3CN and 0.1% formic acid at a
flow rate of 300 nl/min). The nano-LC was coupled online through HDMS Synapt G2 mass spectrometer (Waters Corporation). The data was acquired using Masslynx version 4.1 software (Waters Corporation) in a positive ion mode. LC
–MS data were collected using data-independent acquisition mode MSE with online ion mobility separation. Mass range was set to 50
–2,000 Da with a scan time set to 0.75 s. A reference compound [Glu1]-Fibrinopeptide B (Waters Corporation) was continu- ously infused (500 fmol/ml at a
flow rate 500 nl/min) and scanned every 30 s for online mass spectrometer calibration purpose.
Data processing, searching, and analysis
Raw data
files were processed and searched using ProteinLynx Global SERVER (PLGS) version 3.0.1 (Waters Corporation). Mycobacterium protein sequence database from uniprot (September 29, 2017) was used. The following parameters were used to generate peak lists: (1) minimum intensity for precursors
408 KORABLIOVIENĖET AL.
Acta Microbiologica et Immunologica Hungarica 65, 2018
was set to 135 counts, (2) minimum intensity for fragment ions was set to 25 counts, and (3) intensity was set to 750 counts. Processed data were analyzed using trypsin as the cleavage protease, one missed cleavage was allowed,
fixed modi
fication was set to carbamidomethylation of cysteines, and variable modi
fication was set to oxidation of methionine. Minimum identi
fication criteria included one fragment ions per peptide, three fragment ions per protein and minimum of two peptides per protein. The false discovery rate (FDR) for peptide and protein identi
fication was determined based on the search of a reversed database, which was automatically generated using PLGS when global FDR was set to 4%.
Statistical analysis
Statistically signi
ficant differences between the groups were examined by the Mann
–Whitney U test and Wilcoxon test; p
<0.05 was considered statistically signi
ficant, p
<0.09
–clear trend.
Results
GEP role in bacterial growth
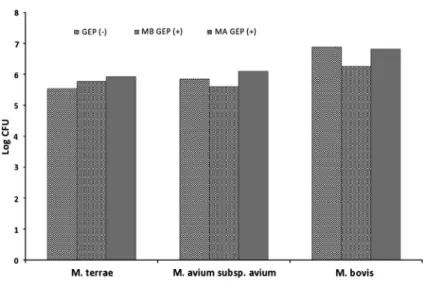
We found that both GEPs inhibited the growth of M. bovis in vitro (Figure
1).The strongest inhibitory process was observed in M. bovis affected with MB GEP (p
=0.030). Our results indicated that MA GEP stimulated the growth of the M.
avium subsp. avium, whereas MB GEP inhibited the process (Figure
1). BothGEPs stimulated the growth of the M. terrae. The strongest stimulation process was observed in M. terrae affected with MA GEP (Figure
1). Statistical signifi- cance of results is given in Table
I.GEP role in bacterial bio
film formation
Cellulose is a critical component of mycobacteria biofilms [9], we scraped
the bio
film biomaterial and stained bio
films cellulose with Congo red. We
observed that higher OD was in samples affected by GEP. OD measurement
and visual analysis con
firm that GEP plays an important role in bio
film formation
process. In samples affected by GEP was enhanced bacterial pellicles, clumps, and
aggregates formation process. The most striking OD and visual changes were in
the M. bovis samples affected by MA GEP (Figure
2). We found thatM. bovis and
M. avium subsp. avium affected by GEP have tendency (p
=0.083) for higher OD.
Statistical significance of results is given in Table
II. We did notfind any statisticallysigni
ficance or tendency in samples with M. terrae.
GEP identi
fication
We analyze GEP samples using FASP method and found 22 proteins. We found 20 proteins in MB GEP and two uncharacterized proteins in MA GEP samples (Table
III). In samples fromM.terrae, we did not
find any proteins that were identi
fiable in uniprot database.
Figure 1.Effect of MB GEP (+) and MA GEP (+) on Log CFU ofM. terrae, M. avium subsp.
avium, andM.bovis.Mycobacterial samples without GEP (−) were considered to be control. GEP:
growth-affecting protein; MA GEP: GEP extracted fromM. avium subsp. avium; MB GEP: GEP extracted fromM. bovis
Table I.Statistical significance of MB GEP and MA GEP on CFU/ml of mycobacteria
Mycobacteria GEP Mean rank Mann–WhitneyU WilcoxonW z p
M. avium subsp. avium
– 10.25 2.500 57.500 −1.623 0.121
MB GEP 5.75
– 3.75 4.500 7.500 −1.186 0.273
MA GEP 7.05
M. bovis – 11.5 0.000 55.000 −2.152 0.030
MB GEP 5.5
– 8.0 7.000 62.000 −0.646 0.606
MA GEP 6.2
Note: Mycobacterial samples without GEP (−) were considered to be control. GEP: growth-affecting protein; MA GEP: GEP extracted fromM. avium subsp. avium; MB GEP: GEP extracted fromM. bovis.
410 KORABLIOVIENĖET AL.
Acta Microbiologica et Immunologica Hungarica 65, 2018
Discussion
T7S and general secretion pathways associated with mycobacteria bio
film formation and growth processes
As mentioned above in samples affected by GEP were enhanced bacterial pellicles, clumps, and aggregates formation. The most striking OD and visual changes were in the M. bovis samples. We found that M. bovis and M. avium subsp. avium affected by GEP has tendency for higher OD.
Figure 2.Effect of MB GEP (+) and MA GEP (+) on biofilms scraped fromM. terrae, M. avium subsp. avium, andM. boviswas evaluated by cellulose optical density (OD) measurement.
Mycobacterial samples without GEP (−) were considered to be control. GEP: growth-affecting protein;
MA GEP: GEP extracted fromM. avium subsp. avium; MB GEP: GEP extracted fromM. bovis
Table II.Statistical significance of MB GEP and MA GEP on optical density of cellulose
Mycobacteria GEP Mean rank Mann–WhitneyU WilcoxonW z p
M. bovis – 1.50 0.000 3.000 −1.732 0.083
MA GEP 4.00
– 1.50 0.000 3.000 −1.732 0.083
MB GEP 4.00
M. avium subsp. avium
– 1.50 0.000 3.000 −1.732 0.083
MB GEP 4.00
– 1.50 0.000 3.000 −1.732 0.083
MA GEP 4.00
Note: Mycobacterial samples without GEP (−) were considered to be control. GEP: growth-affecting protein; MA GEP: GEP extracted fromM. avium subsp. avium; MB GEP: GEP extracted fromM. bovis.
TableIII.MBandMAGEP.First20proteinsweredetectedinMBGEPandtwouncharacterizedproteinsinMAGEP No.ClusternameGenenameStatus*kDaLocationFunctionSuperfamilySecretionpathway 1Majorsecreted immunogenic proteinmpb70 Mpb70Proteinpredicted19Extracellularprotein.Secreted fromthemycobacterialcell. Proteinisabundantly expressedandsecretedinto theculturemedium[12,13]
Geneencode preproteins withsignal peptides[14]
FAS1domainAssociatedwithorareencodedas precursorproteinswithtypical signalpeptidesforexportthrough thegeneralsecretorypathway. Proteinsprocessedbythissystem harboranNterminalsignal peptidethatiscleavedoffasthe proteinisreleasedontheexterior ofthecell.Proteintransport throughtheSecYEG-integral membranecomplex[15,16]
2Cellsurface lipoprotein (fragment)
Mpb83Proteinpredicted20.2Mycobacterialmembrane. Foundintheculturefiltrateof bacteriagrowninliquid culture[17,18] 3Immunogenic protein (fragment) Mpb64Proteinpredicted24.8Associatedwithexocrine proteinfoundintheculture fluid.Secretedprotein associatedwithextracellular region[19,20]
PdaC/RsiV-like 4Membrane proteinB7S04_ 19330Proteinpredicted42.9Associatedwith transmembrane.Integral componentofmembrane
Membraneproteinsinvolvedin thecellenvelope.Associated withenergymetabolic functions[21]
N/AN/A 5ESX-1secretion- associated proteinEspL
B7S05_ 20825Proteininferredfrom homology12.1EsxproteinsofEsx-1are generallysecretedindifferent mediaandnotstrictly regulated[22,23]
Proteins(Esx)lacksignal peptidesandrelyonESX systemsforsecretion[24]
Nucleoid-associatedprotein YbaB/EbfCAssociatedwiththetypeVII secretion(T7S)pathway[25] 6PEfamilyprotein (uncharacterized)B7S04_ 11265Proteinpredicted41.9Associatedwiththe“cellwall andcellprocesses”functional category[26].Detectedin culturefiltrate[27]
Proteinassociated withuntypical signalpeptides[28]
Associatedorbelongtothe samePfamprotein superfamily,designatedthe EsxABclan[29] 7EsxQMb1595_ p3356Proteinpredicted13.0AssociatedwithESAT6Proteins(Esx)lacksignal peptidesandrelyonESX systemsforsecretion8ESAT-6-like protein(fragment)N/AProteininferredfrom homology10.6ESAT6hasbeenreportedtobea cellwallprotein.Secreted, culturefiltrateprotein[30,31]9ESAT-6-like proteinesat6Proteininferredfrom homology9.89 10ATPsynthase subunitalphaatpAProteininferredfrom homology59.2Innermembraneprotein[32]ATPsynthaseis reportedtobe essentialinMycobacteriumfor optimalgrowth[33]
P-loopcontainingnucleoside triphosphatehydrolases 11Conjugaltransfer proteinRN06_ 4459Proteinpredicted22.1N/ACanbeassociatedwithseveral distinctmetabolicprocesses [34]
TypeII/IVsecretionsystemprotein 12ProbableDNA helicaseBCG_ 0913cProteinpredicted59.7Mostofthereplisome componentsareconserved acrossbacteria[35]
Helicasesaremotorenzymesthat separate/unwindduplexnucleic acidstrands[36]
N/A
Putative transferaseBCG_ 1438cProteinpredicted22.8N/AN/AS-adenosyl-L-methionine- dependent methyltransferase
N/A Uncharacterized proteinBCG_ 0394cProteinpredicted22.8N/AN/AThioesterase/thiolester dehydrase-isomeraseN/A Methylated-DNA– protein-cysteine methyltransferase
RN06_ 1638Proteinpredicted5.7N/AInvolvedinthecellulardefense againstthebiologicaleffectsof O6-methylguanineandO4- methylthymineinDNA.Canbe associatedwithDNArestoring afterdamage[37]
MethylatedDNA-protein cysteinemethyltransferase, C-terminaldomain
N/A F420-dependent glucose-6- phosphate dehydrogenase
fgdProteininferredfrom homology37.5Canbedetected incellextractsof mycobacteria[38]
Appearstohavearolein resistancetooxidativestress, viaitsconsumptionofG6Pthat servesasasourceofreducing powertocombatoxidative stressinmycobacteria.
Bacterialluciferase-likePentosephosphatepathway enzyme[39] GTP3′,8-cyclasemoaAProteininferredfrom homology39MoaAislocated onaplasmid[40]Catalyzesthecyclizationof GTPto(8S)-3′,8-cyclo-7,8- dihydroguanosine5′- triphosphate
BelongstotheradicalSAM superfamilyThisproteinisinvolved pathwaymolybdopterin biosynthesis,whichis Cofactorbiosynthesis BacterioferritinbfrAProteininferredfrom homology18.3Ironstoragewithinbacterial cells[41]Iron-storageprotein.Interactive partnersofbacterioferritinand ferritinaredirectlyorindirectly involvedinM.tuberculosis growth,homeostasis,iron assimilation,virulence, resistance,andstresses[42]
Ferritin-likeAssociatedwiththeiron pathways[43] Phenolpthiocerol synthesistype-I polyketide synthaseppsD
ppsDProteinpredicted19.3Inmicrobespolyketidesare frequentlyproducedinculture afteraperiodofactivegrowth hasdepletedthesubstrate [44,45]
Polyketidesynthasesareafamily ofmultidomainenzymesor enzymecomplexesthatproduce polyketidesstructurallydiverse secondarymetabolites,manyof whichhaveantibioticor anticanceractivity,playother rolesintheenvironmentother thantodefeatmicrobial competitors[46,47]
Thiolase-likeType-Ipolyketidesynthase
TableIII.(Cont.) No.ClusternameGenenameStatus*kDaLocationFunctionSuperfamilySecretionpathway 20ThioredoxinB7S05_ 20990Proteininferredfrom homology12.5Cytoplasmic protein[48]ThioredoxinsandTrxRhavebeen showntobeinvolvedin reductionofperoxidesand dinitrobenzenesandalsoto detoxifyhydroperoxidesin vitro[49,50]
Thioredoxin-likeAssociatedwithtypeIIIsecretion system[51] 1Uncharacterized proteinRN06_ 2833Proteinpredicted3.1N/AN/AN/AN/A 2Uncharacterized proteinB7S04_ 00145Proteinpredicted13.1N/AN/AN/AN/A
EsxQ, ESAT-6-like proteins, and ppsD are secreted into the culture medium and can be detectable in culture
filtrate. Mpb70, Mpb83, Mpb64, EspL, PE family and EsxQ, and ESAT-6-like proteins are associated with signal peptides. All these proteins are associated with T7S and general secretion pathways (Table
III).Our results indicate that all GEPs inhibited the growth of the M. bovis. MB GEP inhibited the growth of the M. avium subsp. avium. The strongest inhibitory process was observed in M. bovis affected with MB GEP. As discussed above, mycobacteria differently react to their own and closely related slow-growing organism-secreted proteins. The results suggest that MB GEP inhibited M. bovis growth, while M. avium subsp. avium was stimulated by their own secreted GEP.
There is a lack of information about how mycobacteria responds to their own and closely related, slow-growing organism-secreted proteinaceous compounds.
We identi
fied GEP substrate and found that most of the GEP proteins associated with the T7S pathway. Our
findings suggest that these mycobacteria-secreted GEP are involved in bio
film formation and growth-affecting processes.
The addition of GEP to liquid culture medium should aid the resumption of normal bacteria growth, which could potentially improve the diagnosis and quanti
fication of mycobacterial infections. They may be involved in mycobacterial reactivation. As well as, these proteins can act as inhibitors. Our results contribute to a better understanding of the mechanisms underlying mycobacterial bio
film formation and growth-affecting processes and better characterization of myco- bacterial proteins and their functions.
Acknowledgements
The authors are grateful to Mrs. Rita Viliene for her technical assistance.
Conflict of Interest
The authors declare no con
flict of interest.
References
1. Li, Y. H., Tian, X.: Quorum sensing and bacterial social interactions in biofilms. Sensors 12, 2519–2538 (2012).
2. Waters, C. M., Bassler, B. L.: Quorum sensing: Cell-to-cell communication in bacteria.
Annu Rev Cell Dev Biol21, 319–346 (2005).
3. Sharma, I. M., Petchiappan, A., Chatterji, D.: Quorum sensing and biofilm formation in mycobacteria: Role of C-di-GMP and methods to study this second messenger. IUBMB Life66, 823–834 (2014).
4. Chen, J., Xie, J.: Role and regulation of bacterial LuxR-like regulators. J Cell Biochem112, 2694–2702 (2011).
5. Santos, C. L., Correia-Neves, M., Moradas-Ferreira, P. A., Mendes, M. V.: A walk into the LuxR regulators of actinobacteria: Phylogenomic distribution and functional diversity.
PLoS One7, 46758 (2012).
6. Polkade, A. V., Mantri, S. S., Patwekar, U. J., Jangid, K.: Quorum sensing: An under- explored phenomenon in the phylum Actinobacteria. Front Microbiol7, 131 (2016).
7. Ojha, A. K., Baughn, A. D., Sambandan, D., Hsu, T., Trivelli, X., Guerardel, Y., Alahari, A., Kremer, L., Jacobs, W. R., Jr., Hatfull, G. F.: Growth of Mycobacterium tuberculosisbiofilms containing free mycolic acids and harbouring drug-tolerant bacteria.
Mol Microbiol69, 164–174 (2008).
8. Hans-Curt, F., Wingender, J.: The biofilm matrix. Nat Rev Microbiol 8, 623–633 (2010).
9. Trivedi, A., Mavi, P. S., Bhatt, D., Kumar, A.: Thiol reductive stress induces cellulose- anchored biofilm formation in Mycobacterium tuberculosis. Nat Commun 7, 11392 (2016).
10. Fong, J. N. C., Yildiz, F. H.: Biofilm matrix proteins. Microbiol Spectr3, 10 (2015).
11. Wisniewski, J. R., Zougman, A., Nagaraj, N., Mann M.: Universal sample preparation method for proteome analysis. Nat Methods6, 359–362 (2009).
12. Lounatmaa, K., Brander, E.: Immunoelectron microscopic localization of 22 kDa protein antigen in the surface layer of Mycobacterium bovis BCG strains. In Lounatmaa, K., Brander, E. (eds): Proceedings of the XIIth International Congress for Electron Microscopy.
San Francisco Press Inc., San Francisco, 1990, pp. 894–895.
13. Abou-Zeid, C., Harboe, M., Rook, G. A.: Characterization of the secreted antigens of Mycobacterium bovisBCG: Comparison of the 46-kilodalton dimeric protein with proteins MPB64 and MPB70. Infect Immun55, 3213–3214 (1987).
14. Wiker, H. G., Harboe, M., Nagai, S.: A localization index for distinction between extracellular and intracellular antigens of Mycobacterium tuberculosis. J Gen Microbiol 137, 875–884 (1991).
15. Wiker, H. G.: MPB70 and MPB83– Major antigens of Mycobacterium bovis. Scand J Immunol69, 492–499 (2009).
16. Jungnickel, B., Rapoport, T., Hartmann, E.: Protein translocation: Common themes from bacteria to man. FEBS Lett6, 73–77 (1994).
17. Malen, H., Berven, F. S., Softeland, T., Arntzenn, M. O., D’santos, C. S., De Souza, G. A., Wiker, H. G.: Membrane and membrane-associated proteins in Triton X-114 extracts of Mycobacterium bovis BCG identified using a combination of gel-based and gel-free fractionation strategies. Proteomics8, 1859–1870 (2008).
18. Harboe, M., Wiker, H. G., Ulvund, G., Lund-Pedersen, B., Andersen, A. B., Hewinson, R. G., Nagai, S.: MPB70 and MPB83 as indicators of protein localization in mycobacterial cells. Infect Immun66, 289–296 (1998).
19. Harboe, M., Nagai, S., Patarroyo, M. E., Torres, M. L., Ramirez, C., Cruz, N.: Properties of proteins MPB64, MPB70, and MPB80 of Mycobacterium bovis BCG. Infect Immun 52, 293–302 (1986).
416 KORABLIOVIENĖET AL.
Acta Microbiologica et Immunologica Hungarica 65, 2018
20. Yamaguchi, R., Matsuo, K., Yamazaki, A., Abe, C., Nagai, S., Terasaka, K., Yamada, T.:
Cloning and characterization of the gene for immunogenic protein MPB64 ofMycobacte- rium bovisBCG. Infect Immun57, 283–288 (1989).
21. Gu, S., Chen, J., Dobos, K. M., Bradbury, E. M., Belisle, J. T., Chen, X.: Comprehensive proteomic profiling of the membrane constituents of aMycobacterium tuberculosisstrain.
Mol Cell Proteom2, 1284–1296 (2003).
22. Raman, S., Puyang, X., Cheng, T. Y., Young, D. C., Moody, D. B., Husson, R. N.:
Mycobacterium tuberculosisSigM positively regulates Esx secreted protein and nonribo- somal peptide synthetase genes and down regulates virulence-associated surface lipid synthesis. J Bacteriol24, 8460–8468 (2006).
23. Agarwal, N., Woolwine, S. C., Tyagi, S., Bishai, W. R.: Characterization of theMycobac- terium tuberculosissigma factor SigM by assessment of virulence and identification of SigM-dependent genes. Infect Immun1, 452–461 (2007).
24. Abdallah, A. M., Gey van Pittius, N. C., Champion, P. A., Cox, J., Luirink, J., Vandenbroucke-Grauls, C. M., Appelmelk, B. J., Bitter, W.: Type VII secretion – Mycobacteria show the way. Nat Rev Microbiol5, 883–891 (2007).
25. Houben, N. G., Korotkov, K. V., Bitter, W.: Takefive–Type VII secretion systems of mycobacteria. Biochim Biophys Acta8, 1707–1716 (2014).
26. Mazandu, G. K., Mulder, N. J.: Function prediction and analysis of Mycobacterium tuberculosishypothetical proteins. Int J Mol Sci13, 7283–7302 (2012).
27. Fishbein, S., van Wyk, N., Warren, R. M., Sampson, S. L.: Phylogeny to function: PE/PPE protein evolution and impact onMycobacterium tuberculosispathogenicity. Mol Microbiol 5, 901–916 (2015).
28. de Souza, G. A., Leversen, N. A., Malen, H., Wiker, H. G.: Bacterial proteins with cleaved or uncleaved signal peptides of the general secretory pathway. J Proteomics2, 502–510 (2011).
29. Chen, J. M., Zhang, M., Rybniker, J., Basterra, L., Dhar, N., Tischler, A. D., Pojer, F., Cole, S. T.: Phenotypic profiling of Mycobacterium tuberculosisEspA point-mutants reveals blockage of ESAT-6 and CFP-10 secretion in vitro does not always correlate with attenuation of virulence. J Bacteriol24, 5421–5430 (2013).
30. Majlessi, L., Brodin, P., Brosch, R., Rojas, M. J., Khun, H., Huerre, M., Cole, S. T., Leclerc, C.: Influence of ESAT-6 secretion system 1 (RD1) ofMycobacterium tuberculosis on the interaction between mycobacteria and the host immune system. J Immunol174, 3570–3579 (2005).
31. Stanley, S. A., Raghavan, S., Hwang, W. W., Cox, J. S.: Acute infection and macrophage subversion byMycobacterium tuberculosisrequire a specialized secretion system. Proc Natl Acad Sci U S A22, 13001–13006 (2003).
32. Lai, E. M., Nair, U., Phadke, N. D., Maddock, J. R.: Proteomic screening and identification of differentially distributed membrane proteins inEscherichia coli. Mol Microbiol52, 1029–1044 (2004).
33. Sassetti, C. M., Boyd, D. H., Rubin, E. J.: Genes required for mycobacterial growth defined by high density mutagenesis. Mol Microbiol48, 77–84 (2003).
34. Lagrost, L., Desrumaux, C., Masson, D., Deckert, V., Gambert, P.: Structure and function of the plasma phospholipid transfer protein. Curr Opin Lipidol9, 203–209 (1998).
35. Robinson, A., Causer, R. J., Dixon, N. E.: Architecture and conservation of the bacterial DNA replication machinery, an underexploited drug target. Curr Drug Targets13, 352–372 (2012).
36. Matson, S. W., Kaiser-Rogers, K. A.: DNA helicases. Annu Rev Biochem59, 289–330 (1990).
37. Tuteja, N., Singh, M. B., Misra, M. K., Bhalla, P. L., Tuteja, R.: Molecular mechanisms of DNA damage and repair: Progress in plants. Crit Rev Biochem Mol Biol 36, 337–397 (2001).
38. Purwantini, E., Daniels, L.: Purification of a novel coenzyme F420-dependent glucose- 6-phosphate dehydrogenase from Mycobacterium smegmatis. J Bacteriol10, 2861–2866 (1996).
39. Hasan, M. R., Rahman, M., Jaques, S., Purwantini, E., Daniels, L.: Glucose 6-phosphate accumulation in Mycobacteria: Implications for a novel F420-dependent anti-oxidant defense system. J Biol Chem25, 19135–19144 (2010).
40. Solomon, P. S., Shaw, A. L., Lane, I., Hanson, G. R., Palmer, T., McEwanl, A. G.:
Characterization of a molybdenum cofactor biosynthetic gene cluster in Rhodobacter capsulatuswhich is specific for the biogenesis of dimethylsulfoxide reductase. Microbiol- ogy142, 1421–1429 (1999).
41. Masse, E., Gottesman, S.: A small RNA regulates the expression of genes involved in iron metabolism inEscherichia coli. Proc Natl Acad Sci U S A99, 4620–4625 (2002).
42. Sharma, D., Bisht, D.: Role of bacterioferritin & ferritin inM. tuberculosispathogenesis and drug resistance: A future perspective by interactomic approach. Front Cell Infect Microbiol7, 1–5 (2017).
43. Hameed, S., Pal, R., Fatima, Z.: Iron acquisition mechanisms: Promising target against Mycobacterium tuberculosis. Open Microbiol J9, 91–97 (2015).
44. Bennett, J. W.: From molecular genetics and secondary metabolism to molecular meta- bolites and secondary genetics. Can J Botany73, 917–924 (1995).
45. Bode, H. B., Bethe, B., Höfs, R., Zeeck, A.: Big effects from small changes: Possible ways to explore nature’s chemical diversity. Chembiochem3, 619–627 (2002).
46. Staunton, J., Weissman, K. J.: Polyketide biosynthesis: A millennium review. Nat Prod Rep 18, 380–416 (2001).
47. Adusumilli, S., Mve-Obiang, A., Sparer, T., Meyers, W., Hayman, J., Small, P. L.:
Mycobacterium ulcerans toxic macrolide, mycolactone modulates the host immune response and cellular location of M. ulcerans in vitro and in vivo. Cell Microbiol 7, 1295–1304 (2005).
48. Aslund, F., Beckwith, J.: The thioredoxin superfamily: Redundancy, specificity, and gray- area genomics. J Bacteriol181, 1375–1379 (1999).
49. Zhang, Z., Hillas, P. J., Ortiz de Montellano, P. R.: Reduction of peroxides and dini- trobenzenes byMycobacterium tuberculosisthioredoxin and thioredoxin reductase. Arch Biochem Biophys363, 19–26 (1999).
50. Jaeger, T., Budde, H., Flohe, L., Menge, U., Singh, M., Trujillo, M., Radi, R.: Multiple thioredoxin-mediated routes to detoxify hydroperoxides inMycobacterium tuberculosis.
Arch Biochem Biophys423, 182–191 (2004).
51. Negrea, A., Bjur, E., Puiac, S., Ygberg, S. E., Åslund, F., Rhen, M.: Thioredoxin 1 participates in the activity of theSalmonella entericaserovar typhimurium pathogenicity island 2 Type III secretion system. J Bacteriol191, 6918–6927 (2009).
418 KORABLIOVIENĖET AL.
Acta Microbiologica et Immunologica Hungarica 65, 2018