Plant Signaling & Behavior
Nitric oxide is associated with strigolactone and karrikin signal transduction in Arabidopsis roots
--Manuscript Draft--
Full Title: Nitric oxide is associated with strigolactone and karrikin signal transduction in Arabidopsis roots
Manuscript Number: KPSB-2020-0275R1
Article Type: Article Addendum
Keywords: Arabidopsis thaliana; karrikin; strigolactone; nitric oxide; S-nitrosoglutathione reductase
Manuscript Classifications: Arabidopsis; cell signaling; fluorescence microscopy; hormonal cross talk; nitric oxide signaling
Abstract: Nitric oxide (NO), strigolactones (SLs) and karrikins (KARs) are bioactive signal molecules controlling root morphology. Our knowledge regarding the signal interplay of NO with SLs or KARs is limited. Our previous results pointed out that there is signal interplay between SL and S-nitrosoglutathione reductase (GSNOR)-mediated NO/S- nitrosothiol (SNO) levels in Arabidopsis . In this addendum, we further prove that the pharmacological increment of SL levels by the application of rac -GR24 decreases GSNOR abundance, while reducing SL synthesis by TIS108 intensifies GSNOR protein level and possibly promotes NO signaling in Arabidopsis . Additionally, we observed that the endogenous NO level in the roots of htl-3 (KAR receptor) and d14 (SL receptor) mutants is significantly higher compared to the wild-type. These results, together with the previous ones, confirm the interplay of GSNOR-regulated NO/SNO signaling with SL- and KAR-dependent signal transduction in Arabidopsis .
Order of Authors: Dóra Oláh
Árpád Molnár Vilmos Soós Zsuzsanna Kolbert
Response to Reviewers: Reviewer 1
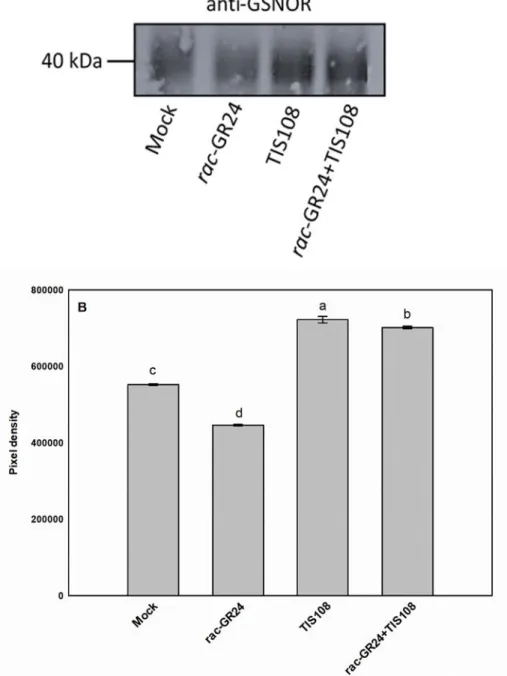
1)Figure 1: The reviewer agrees that GR24 treatment decreased GSNOR protein abundance, while TIS108 treatment increased. When GR24 and TIS108 were applied together, the GSNOR level was very higher than control. The results do not show the complementation test by GR24 treatment. If TIS108 affected as a SL biosynthetic inhibitor, simultaneous treatment of GR24 and TIS108 would decrease by the GSNOR protein levels similar to GR24 treatment. The readers would confuse and could not understand the results. Can TIS108 cancel effects of GR24?
Response: We appreciate this comment. Reviewer 1 is right that co-administration of TIS108 and GR24 increases the amount of GSNOR protein above the control-level and that TIS108 treatment results in a 30% increase compared to control, but TIS + GR24 treatment caused more intense (57%) increase. The strength of the effects may depend on the concentrations. The GR24 and TIS108 concentrations (2 µM and 5 µM, respectively) were chosen based on literature. The applied TIS108 solution were two- times concentrated than GR24 treatment solution which may be the reason why TIS108 had a stronger effect on GSNOR protein than GR24. We think that TIS108 is able to cancel the effect of GR24, as the GR24-triggered GSNOR protein decrease was reversed by TIS108, thus the two treatments may have opposite effects on SL level and also on GSNOR protein abundance.
2) Figure 2: NO levels of htl-3 and d14 are higher than that of Col-0, but NO levels of max2 did not show significant difference compared with Col-0. NO levels of max2 would increase if MAX2 really integrates SL and KAR signaling in the NO production.
Response: Indeed, the data presented here don’t show statistically significant differences, although the averages of max1-1 and max2-1 NO levels are higher by 50 and 60% compared to the WT, respectively (pixel intensities: wild type =528.49,max1- 1=791.48, max2-2= 848.68). In our recent paper (Oláh et al. 2020 Front Plant Sci, 11:1019), the tendencies of NO levels are similar and the mutants’ NO levels are
Powered by Editorial Manager® and ProduXion Manager® from Aries Systems Corporation
significantly higher than the WT. Thus, it can be concluded that max mutants show slightly elevated NO levels compared to WT, and the differences may be or may not be statistically significant depending on how many and which individuals are examined.
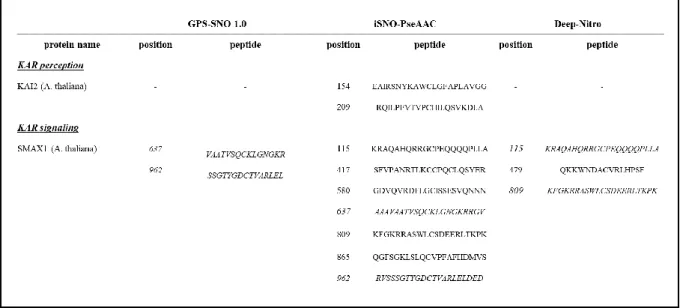
3) Table 1: The authors provided peptide sequences of KAI2 and SMAX1 to evaluate interaction between KAR and NO signal pathways. However, they did not directly evaluate NO-dependent S-nitrosation.
Response: Prior to laboratory experiments, we aimed to predict whether NO may have a direct regulatory role on KAR signaling proteins. Similar in silico prediction was performed in case of proteins involved in SL synthesis and signaling (Kolbert 2019 Physiol Plant, 165:487-497).
4) Do the authors measure SNO levels in the experiments?
Response: Unfortunately, SNO levels have not been measured in htl-3 and d14 mutants yet, but based on previous observations, we hypothesize that they may have higher NO levels compared to WT. In the near future, we would definitely like to measure the SNO levels of the mutants in the frame of a collaboration.
5)There are so many mistakes in the SL-related references. For example, in L.28-30 the authors cited two Nature papers by Umehara et al. and Gomez-Roldan et al. But D27 is not discussed in the papers. DR27 should be revised to D27. L.31-32, Alder et al. and Seto et al. describe that CL is a SL precursor. Booker et al. and Kohlen et al.
did not show what CL is in the papers. L. 32-34, D14 is associated with the SL perception, not MAX2. Reference No. 6 and 16 are the same paper. And others... The authors should carefully consider the reference papers to be used in the text.
Response: We apologize for the mistakes and thank you for drawing our attention to those. We've reviewed and improved the references in the text and also in the list.
Reviewer 2
1. DAF is no longer considered a valid fluorescent probe for authentic nitric oxide visualization.
Please see the references cited below for information on a nitric oxide-specific probe that provides technically more accurate estimation of NO levels.
a) Jain P, David A and Bhatla SC (2016) A novel protocol for detection of nitric oxide in plants. In: Gupta K. (Ed.) Plant Nitric Oxide. Methods in Molecular Biology, vol 1424, pp. 69-79, Humana Press, New York, USA.
b) Keisham M, Jain P, Singh N, von Toerne C, Bhatla SC and Lindermayr C (2019) Deciphering the nitric oxide, cyanide and iron-mediated actions of sodium nitroprusside in cotyledons of salt stressed sunflower seedlings. Nitric Oxide 88: 10-26.
c). Yadav S, David A, Baluška F and Bhatla SC (2013) Rapid auxin-induced nitric oxide accumulation and subsequent tyrosine nitration of proteins during adventitious root formation in sunflower hypocotyls. Plant Signal. Behav. 8: e23196.
d. David A, Yadav S, Baluska F and Bhatla SC (2015) Nitric oxide accumulation and protein tyrosine nitration as a rapid and long distance signaling response to salt stress in sunflower seedlings. Nitric Oxide 50: 28-37.
Response: Although doubts have been raised about the DAF-based NO detection, it is still used in recent, high impact publications (Hartmann et al. 2019, Nature Comm, https://www.nature.com/articles/s41467-019-12045-4; Solhaug et al. 2020, The Plant Journal, https://onlinelibrary.wiley.com/doi/abs/10.1111/tpj.15055; Ding et al. 2020, PNAS, https://www.pnas.org/content/117/20/11147.short etc.).
To support the NO-detecting ability of DAF fluorophore, we incubated Arabidopsis seedlings in Tris-HCl buffer (10 mM, pH7.4; control), in 500 µM SNP solution (in Tris- HCl buffer, diluted from 10mM stock in DMSO) or 500 µM SNP plus 500 µM cPTIO (in Tris-HCl buffer, diluted from 10 mM stock in DMSO) before DAF staining for 60 min under white light illumination. This was followed by a washing step (with Tris-HCl) and the incubation in DAF-FM DA (in Tris-HCl buffer, 10 µM diluted from 5 mM stock in DMSO) for 30 min, room temperature in darkness. We washed the seedlings two-times with buffer and prepared microscopic slides. Data show that SNP treatment
(903.09±44.5) increased DAF fluorescence in the roots of wild-type Arabidopsis compared to control (592.44 ±28.6) while cPTIO application (621.30±48.6) reversed the NO-inducing effect of SNP. These indicate that the intensity of DAF fluorescence changes depending on the NO content of the plant tissue suggesting that the
fluorophore is suitable for detecting NO levels in this experimental system. Similar data supplemented by in vitro measurements, have been reported by Kolbert et al. 2012
2. The observations made using the various mutants of KAR and SL signaling and synthesis are too brief to implicate NO in the signaling routes of KAR and SLs. Similar estimations of GSNOR levels should also have been made in the above cited mutants to authenticate the result as to how the NO/S-nitrosothiol (SNO) levels are affected as a function of enzyme abundance. Moreover, the abundance of GSNOR has been estimated for the whole tissue of wild type plants. More often than not, effects of various molecules are regulated in a tissue-specific manner so it would have been more meaningful to undertake abundance prediction in the roots to provide a better picture of the involvement of modulation of NO/SNO levels as a result of altered GSNOR levels in modeling root architecture.
Response: We agree that GSNOR protein abundance should also be determined in the roots of d14 and htl-3 mutants. The production of sufficient amount of root material in the case of 7-day-old seedlings is time-consuming as well as to perform the
experiment in more biological replicates. Or experiments in the near future will focus on this issue.
3) Another point is that SL-deficient mutants max1-1 and max2-1 exhibit elevated NO/SNO levels due to decrease in the abundance and activity of GSNOR enzyme.
However, in wild type plants exogenously treated with SL mimic, there is a similar decrease in the GSNOR protein abundance. These results seem contradictory as both SL deficiency and exogenous supply of SL elicit a similar response, and thus warrant clarification in the paper.
Response: The lower GSNOR abundance of SL-deficient mutants indeed contradict the GSNOR-decreasing effect of external SL administration. One possible explanation for this may be that the effect of the MAX1 and MAX2 mutations on GSNOR is not SL- specific, but there is some indirect relationship between gene defect and GSNOR protein levels. This possibility was also mentioned in the text, thanks for the suggestion.
L 73-75: “Another possible explanation may be that the effect of the MAX1 and MAX2 mutations on GSNOR is not SL-specific, but there is some indirect relationship between gene defect and GSNOR protein levels.”
Do the levels of NO/SNO also correlate with GSNOR levels determined for the treatments using SL mimic and SL inhibitor? Were there any consequent changes in root architecture?
Response: GR24 treatment reduced GSNOR abundance as well as caused higher NO levels compared to the control (Kolbert 2019 Physiol Plant, 165:487-497), thus there may be an association between the changes caused by GR24. In case of TIS108 treatment, the increase in GSNOR abundance was accompanied by unaffected NO levels. However, it should be noted that the relationship between GSNOR and NO is not direct. The root elongation is oppositely regulated by GR24 and TIS108 (Oláh et al.
2020 Front Plant Sci, 11:1019).
4)Since the three players, NO, KAR and SL, have been implicated in the process of modeling root architecture, were any morphological observations made on roots of the various mutants used in the paper to support the proposition?
Response: Lateral root number has not been determined yet, but primary root length of htl-3, d14 was similar to that of the WT.
We appreciate the useful comments and constructive criticism of the Reviewers. Major modifications are highlighted in yellow in the manuscript file. We are confident about its positive re-evaluation.
Powered by Editorial Manager® and ProduXion Manager® from Aries Systems Corporation
Nitric oxide is associated with strigolactone and karrikin signal transduction in 1
Arabidopsis roots 2
Dóra Oláh
1,2Árpád Molnár
1, Vilmos Soós
3, Zsuzsanna Kolbert
13
4
1
Department of Plant Biology, University of Szeged, Közép fasor 52, H-6726, Szeged, Hungary 5
2
Doctoral School in Biology, Faculty of Science and Informatics, University of Szeged, Szeged, 6
Hungary 7
3
Argicultural Institute, Center for Argicultural Research, Hungarian Academy of Sciences, 8
Martonvásár, Hungary 9
10
Keywords: Arabidopsis thaliana, karrikin, strigolactone, nitric oxide, S-nitrosoglutathione 11
reductase 12
13
Nitric oxide (NO), strigolactones (SLs) and karrikins (KARs) are bioactive signal 14
molecules controlling root morphology. Our knowledge regarding the signal interplay 15
of NO with SLs or KARs is limited. Our previous results pointed out that there is signal 16
interplay between SL and S-nitrosoglutathione reductase (GSNOR)-mediated NO/S- 17
nitrosothiol (SNO) levels in Arabidopsis. In this addendum, we further prove that the 18
pharmacological increment of SL levels by the application of rac-GR24 decreases 19
GSNOR abundance, while reducing SL synthesis by TIS108 intensifies GSNOR 20
protein level and possibly promotes NO signaling in Arabidopsis. Additionally, we 21
observed that the endogenous NO level in the roots of htl-3 (KAR receptor) and d14 22
(SL receptor) mutants is significantly higher compared to the wild-type. These results, 23
together with the previous ones, confirm the interplay of GSNOR-regulated NO/SNO 24
signaling with SL- and KAR-dependent signal transduction in Arabidopsis.
25 26
SLs are recognised as terpenoid lactons consisting of triciyclic lacton (ABC ring) 27
and butenolide group (D ring).
1-4SLs are synthesized from all-trans β-carotene by β- 28
carotene isomerase DWARF 27 (D27) and two carotenoid cleavage dioxygenases 29
CCD7 and CCD8.
5These enzymes produce carlactone (CL) in two subsequent steps 30
from 9-cis-β-carotene. CL is a bioactive precursor of SLs which is oxidized by MORE 31
AXILLARY GROWTH 1 (MAX1)
5yielding canonical and non-canonical SLs. The 32
perception and signal transduction of SL is catalyzed by DWARF14 (D14) and MORE 33
AXILLARY GROWTH 2 (MAX2), respectively.
6D14 is an α/β hydrolase hormone 34
receptor, consisting of a conserved Ser-His-Asp triad.
7D14 binds the hormone, after 35
Olah et al. manuscript R1