CHAPTER 7
Molecular Aspects of the Gene: Replication Mechanisms
ROBERT L . BALDWIN
I . I n t r o d u c t i o n 327 I I . S t r u c t u r e a n d C h e m i s t r y of t h e G e n e t i c M a t e r i a l 328
A . C o v a l e n t S t r u c t u r e of D N A a n d R N A 328 B . P h y s i c a l S t r u c t u r e of t h e D N A H e l i x 332 C . B a c t e r i a l a n d Viral C h r o m o s o m e s 339
I I I . R e p l i c a t i o n of D N A 343 A . E n z y m i c S y n t h e s i s of D N A 343
B . S e m i c o n s e r v a t i v e R e p l i c a t i o n 348
C . S e q u e n t i a l S y n t h e s i s 356 D . C u r r e n t P r o b l e m s 358 E . C o m p a r i s o n w i t h t h e D N A - D i r e c t e d S y n t h e s i s of R N A 364
R e f e r e n c e s 367 I. Introduction
I n the last t e n years we have seen the beginning and rapid growth of research aimed at describing in purely chemical terms a basic biological problem, replication of the genes. T h e problem has two p a r t s : determina
tion of the chemical structures of genes and chromosomes, and definition of the reactions responsible for their replication. A major goal is to repro
duce these reactions in the test t u b e : to isolate the necessary enzymes, find conditions where they function properly, and finally to synthesize in vitro biologically active nucleic acids.
Another major goal is to determine the mechanism of gene replication.
The outlines of the problem are clear b u t an intensive study of mechanism is only beginning. I n general terms the problem of how D N A (deoxyribo
nucleic acid) is replicated was analyzed correctly by Watson and Crick1 on the basis of their structural model for D N A . Their proposal t h a t each polynucleotide chain of the double helix serves as a template for fashioning a new chain of complementary base sequence has been confirmed by t h e enzymic studies of Kornberg and his co-workers (for a summary, see Kornberg2). Detailed studies of the mechanism of D N A synthesis have had to wait upon enzyme purification. Recently the D N A synthesizing enzyme from Escherichia coli has been purified to a stage approximating final purity and D N A polymerases from other sources have been highly purified. T h e enzymes necessary for synthesis of viral R N A ' s (ribonucleic acid) are being purified as this is written,* and R N A polymerases catalyzing the
* Note added in proof: S e e , for e x a m p l e , t h e s e c t i o n on " T h e S y n t h e s i s and Struc
ture of R N A " in Cold Spring Harbor Symposia Quant. Biol. 28, 59-109 (1963).
327
328 ROBERT L. BALDWIN
synthesis of specific R N A under the direction of a D N A template have been purified from several sources.
The purpose of this chapter is to discuss current problems, rather t h a n to provide a complete survey of the literature. (Often a single reference will be given to some point although several workers have contributed t o establishing it, and the basis for choosing this reference m a y be only t h a t it contains a good summary of the evidence.) For other recent reviews, see the articles by Bessman2 a and Bollum.2 b
II. Structure a n d Chemistry o f the G e n e t i c M a t e r i a l
So far as is known now, bacterial and viral chromosomes are large molecules of nucleic acid. We begin, then, with the structure and chemistry of D N A and R N A and consider those aspects which are particularly rele
v a n t to the mechanism of replication. As compared to proteins, t h e struc
tures of nucleic acids are characterized by great simplicity. However, the same terms needed to describe different levels of structural organization in proteins3 are often applied to nucleic acids: these are primary, secondary, and tertiary structure. T h e primary structure is the covalent structure:
the sequence of units in a polynucleotide chain, and the covalent bonds which link these units. Both R N A and D N A have been found to have simple linear structures, without branching. T h e secondary structure is the local conformation in which a nucleic acid is folded or coiled. For most D N A ' s this conformation is known to be the double helix proposed by Watson and Crick4 which is formed from two D N A chains running in opposite directions. T h e tertiary structure is t h e arrangement in three- dimensional space of the entire molecule. Little is known about the tertiary structures of R N A and D N A b u t it is commonly assumed t h a t , unlike proteins, most nucleic acids do not have fixed tertiary structures in solu
tion b u t rather are displayed in space as randomly as the primary and secondary structures will allow. When nucleic acids are assembled into viruses and bacteriophages, regular packing of the nucleic acid moiety is very likely. F o r example, in coliphage T 2 a D N A double helix whose extended length is about 50-70 μ is packed into a phage head whose diam
eter is only 0.1 μ.5
A. COVALENT STRUCTURE OF D N A AND R N A 1. PRIMARY STRUCTURE IN OUTLINE
The covalent structure of a D N A chain is shown diagrammatically in Fig. 1. T h e R N A chain differs only in t h a t ribose replaces deoxyribose.
For a recent summary of bond angles, bond distances, and proof of struc
ture, see Steiner and Beers.6 T h e positions of the glycosidic (base-sugar)
7. MECHANISMS OF G E N E REPLICATION 329 linkages have been confirmed by synthesis and also the glycosidic linkage has been shown to be β. Chemical evidence t h a t the phosphate linkage proceeds from the 3'-OH of one pentose ring to the 5'-OH of its neighbor (reviewed by Brown and Todd7) is confirmed by the results of enzymic hydrolysis of D N A and R N A to mononucleotides. Digestion either of R N A or D N A by venom diesterase gives nearly a quantitative yield of 5'-P mononucleotides (a nucleotide t h a t has phosphate esterified to the 5'-hydroxyl group) while spleen diesterase, in conjunction with micrococcal endonuclease, gives complete conversion to 3'-P mononucleotides.8 - 1 0
T h e purine and pyrimidine bases which are commonly found in D N A
F I G . 1 . C o v a l e n t s t r u c t u r e of D N A : a s e c t i o n of a p o l y n u c l e o t i d e c h a i n c o n t a i n ing t w o b a s e s , C a n d G. T h i s is an a r t i s t ' s s k e t c h of a brass rod m o d e l , b u i l t t o s h o w t h e b o n d a n g l e s a n d b o n d d i s t a n c e s . T h e h y d r o g e n a t o m s h a v e b e e n o m i t t e d a n d m o s t of t h e c a r b o n a t o m s are n o t l a b e l e d .
and viral R N A are shown in Fig. 2. Guanine, cytosine, and adenine are present both in R N A and D N A , while uracil usually occurs in R N A and thymine (5-methyluracil) is found in D N A . T h e names of these bases will be abbreviated to G, C, A, U, and T, respectively, in the rest of this chapter.
Other bases are also present in certain cases. I n the D N A ' s of coliphages T2, T4, and T6, 5-hydroxymethylcytosine completely replaces cytosine1 1 and is found chiefly in the form of mono- and diglucosylated derivatives (for their structures, see Kuno and Lehman1 2).
Two Bacillus subtilis phages have partially glucosylated D N A ' s in which thymine is replaced by uracil and 5-hydroxymethyluracil, respectively.1 3 1 5 Recently, Gold et al.16 have found DNA-methylating enzymes which are distributed ubiquitously and which convert cytosine to 5-methylcytosine
Ν
330 ROBERT L. BALDWIN
and adenine to 6-methylaminopurine. T h e base compositions of R N A and D N A from various sources are discussed b y Sueoka in Chapter 9 of this book.
N u c l e o s i d e
cyti
u r i d thyn
a d e
g u a
p H 2 p H 7 p H 12
cyti
u r i d thyn
a d e
g u a d i n e
Y f
H ^ N ^ O 1 R
* known structure ' postulaied faufomcr
R ! k
cyti
u r i d thyn
a d e
g u a
inc ( R ' » H ) nldlnc ( R ' « C H3)
Y l
H!
Ύ 1R 1 R 1
R
cyti
u r i d thyn
a d e
g u a
l o s i n e
R R I R
cyti
u r i d thyn
a d e
g u a n o s i n e
Λ R j ή R
0
î» R
F I G . 2 . P r o t o n a t e d forms of t h e b a s e s in R N A a n d D N A ; R = ribose a n d d e o x y - ribose, r e s p e c t i v e l y . Other r e s o n a n c e forms, in w h i c h t h e charges a n d t h e d o u b l e b o n d s o c c u p y other p o s i t i o n s , also c o n t r i b u t e b u t t h e p r o t o n s are l o c a t e d in fixed p o s i t i o n s , as s h o w n h e r e . T h e s t r u c t u r e for A at p H 2 is t a k e n from C o c h r a n ,1 7 7 t h o s e for A , U , a n d Τ a t p H 7 a n d 1 2 from M i l e s ,1 7 8 - 1 7 9 a n d t h o s e for C1 8 0> 1 8 0 a a n d G1 8 1 from M i l e s a n d c o - w o r k e r s .
2. LOCATION OF PROTONS
The mechanism of base pairing in D N A and R N A by specific hydrogen bonding hinges on the location of the ionizable protons in the bases A, G, C, T, and U (see Fig. 4). When Watson and Crick4 proposed t h e pairing of A with Τ and G with C in the D N A helix, it was probable b u t not certain
7. MECHANISMS OF GENE REPLICATION 331 t h a t the four bases were present in their amino and keto, rather t h a n imino and enol, forms. (In reference books of t h a t period G was often written as the enol tautomer.) Since then infrared studies of model com
pounds in D20 (especially the work of Miles with analogs in which a methyl group replaces the proton) have established the existence at neutral p H of the keto and amino forms shown in Fig. 2. In view of the suggestion by Watson and Crick1 7 t h a t spontaneous mutations may arise through mispairing of a base in its rare tautomeric form (cf. Fig. 4c), several methods have been tried for detecting these tautomers. Because the proportion which is in the rare tautomeric form is very small, it has been difficult to measure directly. However, Katritzky and W a r i n g1 7 a estimate, on the basis of acid-base properties of uracil analogs, t h a t the frequency of the enol tautomer of iV-methyluracil is about 10~3 to 10~4, while in the cor
responding 5-bromo compound it is some 10 times higher. Similar calcula
tions1^ for cytosine analogs1 7 b allow the frequency of the imino tautomer of iV-methylcytosine to be estimated at about 10~5. These frequencies are much higher t h a n some observed1 8 rates of spontaneous m u t a t i o n (10~7 — 1 0- 8) which m a y correspond to a transition from an A T to a G C base pair, or vice versa. For a more detailed comparison, one needs to know the reac
tion rates for interconversion of the tautomeric forms, as well as for the appropriate steps in the D N A polymerase reaction.
Near p H 4 protons are added to A, G, and C and near p H 10 protons are removed from G, T, and U. T h e structures which result are also shown in Fig. 2 (for a summary of p Ka values, see Steiner and Beers,6 p . 25).
The phosphate group in the backbone of a polynucleotide chain is a strong acid: the p Ka for the corresponding ionization in mononucleotides is below p H l .6
3. CHEMICAL STABILITY
Depurination and diester bond breakage are the chief types of chemical damage suffered by D N A and R N A . I n D N A strand breakage appears to follow depurination (see the discussion by Fiers and Sinsheimer1 9), and depurination is catalyzed by acid and by h e a t .2 0 I n the case of R N A , diester bond cleavage proceeds directly via a cyclic intermediate involving the 2'-OH group of ribose, and the reaction is strongly catalyzed by alkali.
Alkaline hydrolysis a t room temperature can give quantitative breakdown to mononucleotides. A thorough study of the rates of hydrolysis of R N A under different conditions has been made by Bacher and K a u z m a n n .2 1 One may summarize by saying t h a t R N A is unstable in acid and very unstable in alkali, whereas D N A is unstable in acid and fairly stable in alkali. Both become less stable at any p H as the temperature is increased. With mate
rials isolated from natural sources or made enzymically, enzymic hydrolysis is often a problem.
332 ROBERT L. BALDWIN
Other types of chemical damage have also been demonstrated. For example, ultraviolet light promotes the hydration and dimerization of thymine (reviewed by Wierzchowski and Shugar2 2). Mechanical shear evidently will break covalent bonds in D N A if the molecule is sufficiently large (see Section I I , C ) .
B . PHYSICAL STRUCTURE OF THE D N A HE L I X 1. DERIVATION OF THE STRUCTURE BY X - RA Y DIFFRACTION
T h e structure of the D N A double helix (form B , the one believed to exist in solution) is shown in Fig. 3. Its principal features are well known.
I t has the form of a rope ladder twisted into a regular right-handed helix, in which the base pairs A T and G C are the rungs of the ladder and the sugar-phosphate chains are the two ropes. T h e sugar-phosphate backbone has polarity, since a phosphate connects the 3 '-OH of one deoxyribose to the 5'-OH of the next; the two complementary strands have opposite polarities. There are ten base pairs per t u r n of the helix and they are spaced 3.4 A. apart, the v a n der WaaFs contact distance between stacked purine and pyrimidine rings. T h e A T and GC base pairs have the structures shown in Fig. 4. Each base pair is rotated 36° about the helix axis from its neighbor. An important property of the structure is t h a t the helix is regular : every base-sugar linkage is found at the same distance from the helix axis and is rotated, relative to its neighbors in the chain, by the same amount (36°). The ionized phosphate groups are on the outside of the helix, where they nevertheless produce a large electrostatic charge which tends to unfold the helix. Location of the water molecules around the helix is not shown.
I n fact, it is not known whether they occupy definite positions on the helix.
Water probably plays a role in stabilizing the D N A structure since there is a change to another crystal form, the A form, when wet fibers of sodium D N A are taken to low humidity.2 3
Evidence for the structure shown in Fig. 3 comes both from X - r a y diffrac
tion patterns of D N A fibers and from model building. A chief purpose of building models is to exclude structures which require unusual bond angles or bond distances, or which necessitate placing two nonbonded atoms closer t h a n the sum of their v a n der WaaFs contact distances. Then, when the atomic coordinates of the model have been measured, atomic scattering factors are used in programing a computer to find the X - r a y p a t t e r n which this structure would give. Comparison with the observed p a t t e r n then shows whether or not further adjustments of the model are needed. This was the procedure used by Langridge et a L ,2 4 , 2 5 whose third model gives good prediction of the observed X - r a y patterns. I t retains the essential features of the original model proposed by Crick and W a t s o n2 6 b u t the
7. MECHANISMS OF G E N E REPLICATION 333
F I G . 3 . Space-filling m o d e l of t h e D N A d o u b l e h e l i x , f o r m Β ( m o d e l e d after t h e figure s h o w n b y E i g e n1 8 2 a n d t h e earlier figure of F e u g h e l m a n et al.183).
334 ROBERT L. BALDWIN
coordinates of the atoms have been altered appreciably. The present struc
ture is believed to be a close approximation to the final one.2 5
Three different crystal forms of D N A (the A, B, and C forms) have been
a. Hydrogen bonding of adenine to thymine
b. Hydrogen bonding of guanine to cytosine
c, Possible pairing of guanine to S-bromouracil (enol tautomer)
Ν
A
11\ if V H - ^
ν
N - C / //
\ i ^ C ^ ^ 0
F I G . 4. H y d r o g e n b o n d i n g in t h e A T a n d G C b a s e pairs (a, b) a n d a n e x a m p l e (c) of t h e m i s p a i r i n g t h a t m i g h t occur w h e n one b a s e is p r e s e n t in a rare t a u t o m e r i c f o r m : t h e p o s s i b l e pairing of G w i t h t h e enol t a u t o m e r of 5-bromouracil.
reported in which there are obvious differences in the structure of the helix.
Evidence t h a t the Β form is the one present in solution rests chiefly on finding the Β form at high humidities when the D N A double helices are separated from each other by large sheaths of water.2 3 Also the mass per
7. MECHANISMS OF GENE REPLICATION 335
unit length of D N A in solution has been measured by low-angle X-ray scattering2 7 and agrees with the value predicted for the Β form. A transition to the A form is observed in fibers of sodium D N A when the relative humidity is lowered below about 75 %. Although a complete structure for the A form has not yet been reported, the work of Langridge et al.28 con
firms the suggestion of Crick and W a t s o n2 6 t h a t the base pairs are tilted relative to the helix axis and displaced sideways, giving rise to a 3 0 % shortening of the helix. Highly crystalline X-ray patterns are obtained readily from fibers of t h e A form of D N A , and consequently much of the early effort directed towards solving the structure of D N A was spent in analyzing these patterns. When Langridge et al28 found t h a t the lithium salt of D N A would crystallize in the Β form, they could t h e n obtain suffi
cient information from the X-ray patterns of the Β form to determine its structure in detail.
T h e C form of D N A is given by fibers of lithium D N A under special conditions of salt and low humidity. These conditions and a proposed struc
ture for the C form have recently been given by Marvin et al29 T h e base pairs are tilted 6° relative to the helix axis and the two bases in a base pair are twisted 5° away from each other.
2. UNFOLDING OF THE D N A HE L I X
The D N A helix is a highly ordered structure in which most of the possible rotations about single bonds have been frozen in place. One would expect from this t h a t the separated polynucleotide chains possess more entropy t h a n the helix and, since this will tend to unfold the helix, t h a t there must be a favorable change in heat content (heat must be given off) on forming the double helix from the separated chains. These expectations have been confirmed by experiment. Recent studies on the formation of the R N A double helix rArU,* by mixing the R N A homopolymers of A and U, give AH = — 8.7 kilocalories per mole of base pairs and AS = —23 entropy units per mole of base p a i r s .3 0 , 3 1 This value of — AH is larger t h a n expected for merely forming two hydrogen bonds in aqueous solution.3 0 Since the rArU homopolymer pair forms a double helix whose structure is similar to t h a t of D N A3 2 these values for AH and AS provide a guide to the behavior of D N A .
Granted t h a t AH and AS both are negative, it follows t h a t AG, the change in free energy for the transition from separated strands to helix, will be
come positive above a given temperature, Tm(AG = AH — TAS; at Tm AG =
* H e r e r is u s e d t o d e n o t e a n R N A c h a i n , d t o d e n o t e a D N A c h a i n ; r A r U s t a n d s for a c o m p l e x f o r m e d f r o m s e p a r a t e c h a i n s of rA a n d r U p o l y m e r s w h i l e t h e R N A c o p o l y m e r w h i c h c o n t a i n s A a l t e r n a t i n g w i t h U will be w r i t t e n r A U a n d r a n d o m c o p o l y m e r s as r A , U .
336 ROBERT L. BALDWIN
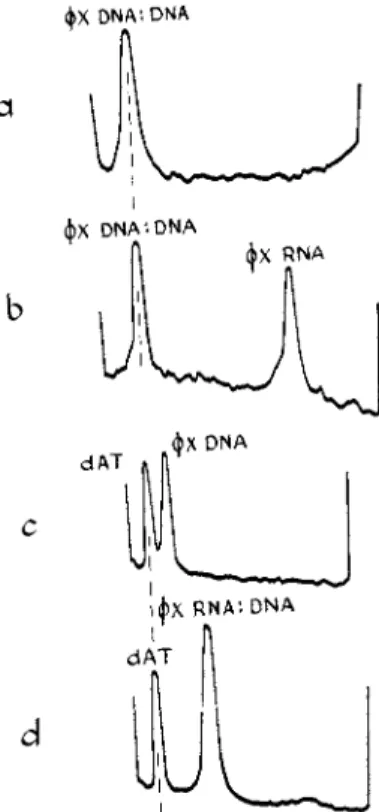
0 and AH = TmAS). Above this temperature the D N A helix will unfold spontaneously. Tm is usually determined from the mid-point of the melting curve of absorbance vs. temperature, with the assumption t h a t the degree of melting is equal to the percent of the final increase in absorbance.3 3 T h e melting temperature of a D N A increases with its GC content3 4 and this has been interpreted in terms of a different AH for A T and GC base pairs.3 5 T h e use of melting curves to compare the stability of different helices, and also to characterize nucleic acids, is illustrated in Fig. 5 .
4 0 5 0 6 0 7 0 8 0 9 0 T e m p e r a t u r e ( * C )
F I G . 5 . M e l t i n g c u r v e s of a b s o r b a n c e v s . t e m p e r a t u r e for t h r e e forms of < £ X - 1 7 4 D N A1 7 4: t h e n a t i v e s i n g l e - s t r a n d e d D N A , t h e R N A : D N A h y b r i d m a d e w i t h t h e R N A p o l y m e r a s e from E. coli, a n d t h e d o u b l e - s t r a n d e d D N A ( D N A : D N A ) m a d e w i t h t h e D N A p o l y m e r a s e from E. coli. T h e s o l v e n t c o n t a i n s N a3 c i t r a t e , 0 . 0 5 M in N a+. A t t h i s N a+ c o n c e n t r a t i o n t h e n a t i v e φΧ. D N A is a l m o s t c o m p l e t e l y m e l t e d at 4 0 ° C , t h e l o w e s t t e m p e r a t u r e s h o w n .6 6
I n addition to the increase in rotational entropy, another major factor tending to unfold the D N A helix is the charge repulsion between ionized phosphate groups, which is less in the separated strands. Consequently the helix can be stabilized* by adding a neutral salt (e.g. NaCl) which screens the phosphate charges or by neutralizing the phosphate groups with a cation such as M g+ + which is bound strongly. The D N A helix also unfolds a t acid or alkaline p H ' s (reviewed by J o r d a n3 6; see also Dove et al.z1). At
* T h i s e x p l a i n s t h e s t r o n g d e p e n d e n c e of Tm on t h e t y p e a n d c o n c e n t r a t i o n of t h e c a t i o n , a n d t h e c o n s e q u e n t u s e of a s t a n d a r d s o l v e n t (e.g., 0 . 1 5 M N a C l , 0 . 0 1 5 M N a 3 c i t r a t e3 4) t o c o m p a r e t h e s t a b i l i t y of different D N A ' s .
7. MECHANISMS OF G E N E REPLICATION 337 these p H ' s , * certain hydrogen bonds between base pairs are broken by the addition or removal of protons; compare Figs. 2 and 4.
Since the D N A helix is formed by winding two chains about each other once every ten base pairs, this helix must be unwound as it is melted. For some time it was thought t h a t the two strands would remain wound around each other after melting.3 8 T h e first clear indication t h a t they do come apart was obtained by Meselson and Stahl,3 9 who studied a hybrid D N A (labeled for one generation of growth with a heavy isotope) and who found t h a t the D N A dissociates into two subunits after melting. These can be separated by centrifugation in a density gradient. Later work has been aimed at finding whether the two subunits are in fact the two strands of the D N A helix (see Section Ι Ι Ι , Β ) .
T h e mechanism of unwinding, and prediction of the time required for unwinding, have been the subject of much theoretical study.4 0"4 3 This problem has a direct bearing on the mechanism of D N A replication4 4 since recent evidence suggests t h a t in m a n y cases the entire D N A content of a bacterial or viral chromosome is a single molecule of nucleic acid (Section II,C). According to the semiconservative model for D N A replication, t h e two strands of a double helix must also be unwound during replication, each strand then becoming part of a new helix. T h e great length of these mole
cules raises certain questions. Given a fixed force which drives the unwind
ing of the helix, how long will the unwinding take? How much work is required to unwind the strands? The problem is t h a t both the time and the energy required increase rapidly with the length of the helix. Depending on the mechanism of unwinding, the time required shows a dependence on helix length L varying from L2 to L3.4 3 There are still few experimental studies in this field, b u t recent work by Freese and Freese4 5 indicates t h a t the time required to unwind the D N A of a bacterial chromosome may be comparable to the generation time (20 minutes or so) if the D N A is a continuous double helix. T h e time required to melt T 2 D N A (molecular weight about 1.3 Χ 107) has been measured by Crothers4 6 and agrees fairly well with the prediction of Fixman4 3 for melting by unwinding from one end of the helix.
When thermally melted D N A is cooled rapidly, or alkali-melted D N A is quickly reneutralized, the D N A remains denatured. The viscosity is then greatly reduced, the molar absorbancy at room temperature is greater t h a n t h a t of native D N A and the absorbancy melting curve is spread out over a wide temperature range (cf. Fig. 12b). Amino groups of A, G, and C are more available for reaction with formaldehyde4 7 or nitrous acid, and
* T h e p H at w h i c h t i t r a t i o n a n d u n f o l d i n g of a D N A occurs is different t h a n t h a t for t i t r a t i o n of t h e m o n o n u c l e o t i d e s , b e c a u s e t h e p r o t o n s are h e l d i n h y d r o g e n b o n d s i n t h e h e l i x .
338 ROBERT L. BALDWIN
the increased rate of reaction with these reagents can be used to distinguish denatured from native D N A . Also certain deoxyribonucleases distinguish native from denatured D N A . T h e best example is E. coli exonuclease I ,4 8·4 9 crystalline preparations of which attack denatured D N A at a rate some 40,000 times greater t h a n native D N A .
Recently Marmur, Doty, and co-workers have made the important dis
covery t h a t the two complementary chains of a D N A helix can recombine after they have been separated by melting.5 0 _ 5 3 a Evidence for specific recombination* is given by recovery of transforming activity, formation of a structure which once more melts sharply at the Tm of the native D N A , and by the formation of hybrid molecules (labeled in one strand by heavy isotopes) which are resistant to breakdown by E. coli exonuclease I. There is not yet evidence t h a t strand recombination can occur in vivo by such a mechanism.
3. SECONDARY STRUCTURE OF R N A AND φ Χ D N A
T h e properties of most viral R N A ' s and also of φ Χ D N A resemble those of denatured D N A more t h a n native D N A and it seems probable t h a t these molecules do not have any fixed and regular secondary structure, although elements of secondary structure are present. For example, the existence of some secondary structure in the R N A from tobacco mosaic virus is shown clearly by the work of Spirin and co-workers5 4 on the viscosity-temperature curves of this R N A . T h e freshly prepared infectious R N A shows a sharp increase in viscosity over a temperature interval of about 10°C. Presumably the formation of some base pairs between A and U and between G and C provides the basis for the structure maintained at low temperatures. I t has been shown by Fresco and Alberts,5 5 from the analysis of mixing curves of the R N A homopolymer of U with a random copolymer of A and U, t h a t an ordered structure will form with A paired to U and with "looping o u t " of the extraneous U residues from the rA,U copolymer, if its U content is not too large. As yet there is no case known in which an R N A must have a spécifie secondary structure to show its biological activity, although the definite possibility exists t h a t this may be true of amino-acid acceptor R N A ' s .5 6
The X-ray diffraction patterns shown by natural R N A ' s are usually indistinct and have not been of much use in establishing secondary structure until recently. Spencer et al.57 have obtained a fairly crystalline fiber pattern from a preparation containing yeast acceptor R N A and later Langridge and G o m a t o s5 8 found t h a t reovirus R N A , whose solution properties indicate t h a t it has a large amount of secondary structure,5 9 gives a clear X-ray
* T h e procedure u s e d for s t r a n d r e c o m b i n a t i o n is " a n n e a l i n g " at a t e m p e r a t u r e a b o u t 25°C. b e l o w Tm in 0.3 M N a C l , 0.03 M N a3 c i t r a t e .5 3
7. MECHANISMS OF GENE REPLICATION 339 diffraction p a t t e r n of a similar type. Both resemble the crystalline pattern found by D a v i e s6 0 for the R N A homopolymer pair r l r C . T h e major point of interest is t h a t the structure resembles the A form, rather t h a n the Β form, of D N A and although constructed from base pairs of the D N A type, they are tilted by about 10-15° from a perpendicular to the helix axis.5 8
Both electron microscopy6 1 and low angle X-ray scattering from solu
t i o n6 2 indicate t h a t R N A can form a rodlike structure in solution over short distances. T h e melting curves of absorbancy vs. temperature are broad for natural R N A ' s6 3 indicating t h a t melting is much less cooperative t h a n in D N A . Also the absorbancy can be increased by reducing the salt concentration,6 4' 6 5 unlike native D N A whose absorbancy remains quite constant.
T h a t the D N A from phage φΧ-174 is single-stranded has been shown in many w a y s .6 6 I t was suggested by P3 2 suicide results.6 7 T h e most direct demonstration was t h a t the base composition does not obey t h e pairing rule for a Watson-Crick helix, in which A = Τ and G = C on a molar basis. T h e fact t h a t the base composition does not follow the pairing rule means also t h a t φ Χ D N A is not an equal mixture of the two complementary strands, and presumably only one type of strand is present. Other indica
tions of the single-stranded nature of φ Χ D N A are t h a t the absorbancy melting curve is broad and melting begins at low temperatures (cf. Fig. 5), the molar absorbancy at 20°C. is higher t h a n t h a t of a native D N A , and there is a significant rate of reaction with formaldehyde.6 6
C. BACTERIAL AND VIRAL CHROMOSOMES 1. ' ON E CHROMOSOME, ON E NUCLEIC ACID MO L E C U L E "
As techniques have been developed for handling giant D N A molecules, the hypothesis has been greatly strengthened t h a t viral and even bacterial chromosomes* are single molecules. This hypothesis was developed clearly by Levinthal6 8 some years ago. T h e recognition t h a t shearing forces devel
oped in routine laboratory operations can break these giant m o l e c u l e s6 8 , 7 0 clarified earlier contradictory results. Thus, while early studies by P3 2 autoradiography suggested t h a t the D N A from phage T 4 contained one large piece of molecular weight 45 Χ 106 in addition to smaller pieces,7 1 ultracentrifugal a n a l y s e s7 2 , 7 3 seemed to show only small pieces. Later experiments with P3 2 autoradiography, which were designed to prevent shear breakage, showed t h a t the entire D N A contents of the phage (about 130 million molecular weight) are present in a single m o l e c u l e .7 4 , 7 5 T h e
* S i n c e t h e c h r o m o s o m e s of p l a n t a n d a n i m a l cells are h i g h l y c o m p l e x s t r u c t u r e s , as s e e n b y t h e e l e c t r o n m i c r o s c o p e , R i s a n d C h a n d l e r6 9 s u g g e s t t h a t t h e t e r m ''chro
m o s o m e " is i n a p p r o p r i a t e here a n d s h o u l d be r e p l a c e d b y t h e t e r m " g e n o p h o r e . "
340 ROBERT L. BALDWIN
length of the T 2 D N A molecule has also been measured directly by electron microscopy5 and by tritium autoradiography7 6 and these results indicate also t h a t there is one D N A molecule per phage. Similar results have been obtained with other phage D N A ' s , for example φΧ-174,6 6 T 3 ,7 7 and T 7 .7 8 None of these results can exclude, of course, the possibility t h a t some non- D N A material links different sections of the chromosome together.*
These conclusions have been anticipated for several years from genetic studies. All of the genetic markers of E. coli belong to a single linkage group.8 0 Furthermore, this linkage group has physical continuity as shown by interrupted mating experiments: transfer of a chromosome from a n Hfr to an F~ strain can be interrupted by shearing in a Waring Blendor.
T h e number of genetic markers which enter a recipient cell is proportional to the time of mating before interruption (see Chapter 1 by Gross). Simi
larly, the genetic markers of phages such as λ and T 4 have been shown to have single linkage g r o u p s .8 1 , 8 2
2. CIRCULAR CHROMOSOMES
The question of whether bacterial and viral chromosomes are circular is presently being studied in several systems. The genetic m a p of E. coli can be represented by a circle.8 0 On the other hand, Hfr strains of E. coli deliver their chromosomes to F~ cells in a mating experiment as if these chromosomes were linear. Very recently Cairns8 3 -8 4 has been able to extract from E. coli chromosomes which in a few instances are intact and untangled, and to photograph them by tritium autoradiography (Fig. 6). His results suggest t h a t both the Hfr and the F~ strains have chromosomes which are physically circular, and t h a t circularity is maintained during replication.
If this is true it is necessary, in order for the two daughter chromosomes to separate from each other after replication, t h a t the continuity of the D N A double helix be interrupted by a "swivel" where free rotation can occur. The swivel could be simply a break in one D N A strand or some non-DNA material.
Preparations of the single-stranded D N A from φΧ-174 show two com
ponents in sedimentation velocity experiments.6 6 Recent e x p e r i m e n t s1 9 , 8 5 , 8 6
show t h a t only the faster of these (which may be called native φ Χ D N A ) is active in transforming protoplasts. The native φ Χ D N A is also resistant to attack by E. coli exonuclease I, an enzyme which readily attacks single-
* W h e n c o n t r o l l e d shear b r e a k a g e is u s e d as a t o o l for o b t a i n i n g D N A f r a g m e n t s of defined l e n g t h ,7 9 n o w e a k p o i n t s are f o u n d in T 2 D N A . C a l c u l a t i o n s s h o w t h a t t h e s h e a r i n g stress s h o u l d be g r e a t e s t in t h e m i d d l e of t h e m o l e c u l e , a n d it is f o u n d t h a t b r e a k a g e does occur near t h e m i d d l e a n d t h a t m o l e c u l e s b e l o w a critical size r e m a i n u n b r o k e n .
SiÏÏÎ
F I G . 6. A u t o r a d i o g r a p h of t h e c h r o m o s o m e of E. coli K 1 2 H f r ,8 4 l a b e l e d w i t h H3- t h y m i d i n e for t w o g e n e r a t i o n s . T h e scale s h o w s 1 0 0 μ. One can s e e t w o circular s t r u c tures (each 1 1 0 0 μ in circumference) j o i n e d a l o n g a c o m m o n s e g m e n t .
3 4 1
342 R O B E R T L. BALDWIN
stranded D N A b u t which requires a 3'-OH end group.4 8 The first cleavage by a n endonuclease (pancreatic deoxyribonuclease) reduces the sedimenta
tion coefficient of the native D N A to t h a t of the original slower moving component, and enables E. coli exonuclease I to act. T r e a t m e n t with spleen diesterase indicates t h a t native φ Χ also lacks a free 5'-OH end group. These results could mean either t h a t the molecule is circular or t h a t the ends are blocked in some way. The fact t h a t the first cleavage of native D N A by endonuclease gives a physically homogeneous product, as judged by sedi
mentation velocity experiments, argues for the ring structure: a linear molecule should be broken into two pieces of different sizes, if the attack is at random. The exonuclease I cannot hydrolyze the broken ring structure completely and seems to reach a block in the chain. Thus, native φ Χ D N A behaves as if it were a circular molecule containing one unusual linkage. *
The chromosome of phage T 4 is genetically circular.8 7 Annealing experi
ments with pieces broken by shear8 8 indicate t h a t the base sequence is circularly permuted from one molecule to the next. This would be the result expected if a circular chromosome were broken at random either on extrac
tion from the phage or before being packed into the phage. T h e existence of "terminal redundancy" hétérozygotes in T 48 7 a leads Streisinger et al.
to suggest a model for circular permutation in which the physical structure is always linear.
Still a different case is presented by phage λ. Its genetic m a p is linear8 1 and shear breakage of the extracted D N A breaks the genetic m a p roughly in h a l f .8 1 , 8 9 · 9 0 T h u s the genetic m a p and the D N A are colinear. However the ends of λ D N A seem to be sticky. Hershey et al.91 have found a folded form as well as dimers and trimers; Ris and Chandler6 9 have found circular structures by electron microscopy, which can be produced at will9 1 a using conditions established by Hershey et al.91 There appears to be a change in the order of the phage genes when λ D N A is attached to the host chromo
some in the lysogenic s t a t e .9 1 b , c
These examples show t h a t circularity is an important question in con
sidering the structure and replication of bacterial and viral chromosomes, and t h a t it is too soon to generalize.
* Note added in proof: T h e " r e p l i c a t i v e f o r m " of φ Χ D N A (see S e c t i o n I I I , D , 4 ) , w h i c h has t h e p r o p e r t i e s of d o u b l e - s t r a n d e d D N A , has b e e n purified a n d f o u n d t o be circular b y direct e x a m i n a t i o n in t h e e l e c t r o n m i c r o s c o p e [A. K . K l e i n s c h m i d t , A.
B u r t o n , a n d R. L. S i n s h e i m e r , Science 142, 961 (1963)]. A l s o , e l e c t r o n m i c r o s c o p e p i c t u r e s [W. S t o e c k e n i u s , Proc. Natl. Acad. Sci. U.S. 50, 737 (1963)] of t h e D N A from p o l y o m a virus (a s m a l l , animal virus) confirm t h e c o n c l u s i o n — b a s e d on p h y s i c o - c h e m i c a l p r o p e r t i e s — t h a t t h e D N A is d o u b l e - s t r a n d e d and circular [R. D u l b e c c o a n d M . V o g t , Proc. Natl. Acad. Sci. U.S. 50, 236 (1963); R. Weil a n d J. V i n o g r a d , Proc.
Natl. Acad. Sci. U.S. 50, 737 (1963)].
7. MECHANISMS OF GENE REPLICATION 343 III. Replication o f D N A
A . ENZYMIC SYNTHESIS OF D N A 1 . COMPONENTS OF THE REACTION
The components needed for D N A synthesis in vitro a r e9 2 (1) the D N A - synthesizing enzyme ( D N A polymerase), (2) deoxynucleoside triphos
phates of the four bases in D N A , or certain of their analogs, (3) M g+ +, and (4) a template D N A . The typical event in synthesis is the addition of a mononucleotide to the growing end of a D N A chain, which is illustrated
F I G . 7. T h e t y p i c a l e v e n t in D N A s y n t h e s i s in vitro: a d d i t i o n of a n u c l e o t i d e at t h e 3' O H e n d of a D N A s t r a n d , g o v e r n e d b y b a s e pairing of t h e a d d e d n u c l e o t i d e t o t h e c o m p l e m e n t a r y s t r a n d .
schematically in Fig. 7. The chain to which addition occurs is called the primer strand and the complementary one is called the template strand.9 3 The presence of a free 3' - O H group, either at the end of the template strand or on the primer strand, greatly stimulates the reaction9 3 b u t it is not yet known whether it represents an absolute requirement. Reversal of the synthetic reaction by pyrophosphorolysis can be demonstrated9 2 , 9 4 b u t its rate is negligible under the usual synthetic conditions.
Initial efforts to demonstrate D N A synthesis in vitro were hindered by the small amounts of D N A polymerase in the cell and by the action of deoxyribonucleases in cell extracts in breaking down D N A , including any D N A newly synthesized. These same two problems have made the purifica
tion of D N A polymerase a time-consuming project, requiring both large quantities of cells and patience. However the D N A polymerase from E. coli
344 ROBERT L. BALDWIN
has now been purified to a point approaching final p u r i t y .9 4 , 9 5 T h e degree of purification, starting from the initial extract, is about 2000-fold and the molecular weight of the E. coli polymerase is about 100,000.9 4 If the activity per molecule of D N A polymerase is the same in the crude extract as in the purified product (when assayed with the d A T copolymer as template) then there are about 300 molecules of D N A polymerase per cell, assuming9 5 a t h a t the weight of protein per cell times Avogadro's number is 6 Χ 101 0. Other sources from which the enzyme has been highly purified include calf t h y m u s ,9 6 - 9 8 extracts of T2-infected E. coli" and B. subtilis,100 and these enzymes show the same basic requirements for catalyzing D N A synthesis.
0 . 6 ι 1 1 1 1 1 Γ "- I I I
Template'- T h y m u s DNA
Time i n m i n u t e s
F I G . 8. A n e x a m p l e of t h e r e q u i r e m e n t for all four t r i p h o s p h a t e s in D N A s y n t h e s i s .1 0 2 In t h e c o m p l e t e s y s t e m , w i t h calf t h y m u s D N A as t e m p l a t e , s y n t h e s i s b e g i n s p r o m p t l y and since t h e p r o d u c t h a s a h i g h v i s c o s i t y t h e r e a c t i o n c a n be f o l l o w e d v i s c o m e t r i c a l l y . W h e n d A T P is o m i t t e d n o s y n t h e s i s occurs w i t h i n t h e t i m e of t h i s e x p e r i m e n t .
2. EVIDENCE TH A T THE TEMPLATE IS COPIED
Two striking features of the enzymic reaction are the requirements for a template D N A and for the triphosphates of all four deoxynucleosides. The latter is shown in Fig. 8. Both requirements suggest t h a t the template D N A is being copied enzymically. Stronger evidence comes from the observation t h a t the newly synthesized D N A has the same base composition as the template when D N A ' s of widely differing AT contents are used as tem
plates.1 0 1 , 1 0 2 Furthermore, the base composition of the product does not change when the relative concentrations of the four triphosphates are varied widely- A more stringent test of accurate copying is the distribution of dinucleotides or "nearest neighbors" (ApC, GpA, etc.) along the D N A chains. The same nearest neighbor frequencies were found using a sample
7. MECHANISMS OF GENE REPLICATION 345 of calf thymus D N A as with the use of an enzymically synthesized product present after 20-fold replication of this D N A .1 0 3 When φ Χ D N A and double- stranded φ Χ D N A (made enzymically) were used as templates, the nearest neighbor frequencies obeyed the rules predicted from the base ratios for φ Χ D N A (in which A ^ T, G ^ C) and for a double-stranded D N A (A = T, G = C), respectively.1 0 4 Thus it seems definite t h a t the enzymic syn
thesis of D N A in vitro is a reaction in which the base sequence of the tem
plate D N A is copied.
Less is known about the accuracy of copying at the macromolecular level. One would like to have the answers to the following questions. (1) Does the D N A polymerase produce in vitro a new complementary strand of exactly the same length as the template strand? (β) Is the newly syn
thesized strand an accurate copy of the template strand along its entire length? These are closely related to a third, more basic question. (3) Can the D N A polymerase initiate new chains, or must the in vitro reaction always proceed by addition to a 3'-OH group on the primer strand? Clearly, unless new chains can be started one cannot expect accurate copying of the length of a template strand. At present definite answers to these questions cannot be given; however, evidence which bears on them will be considered later in the chapter.
3. De Novo SYNTHESIS OF D N A POLYMERS
When the D N A polymerase from E. coli is incubated without a template D N A but in the presence of the triphosphates of A and T, or of G and C, there is no chemically measurable synthesis for several hours and then new polymers are made whose synthesis is rapid once it begins. T h e d A T co
polymer1 0 5 contains A and Τ in strictly alternating sequence while the d G d C homopolymer p a i r1 0 6 contains complementary chains of G and of C.
Synthesis begins promptly when the d A T copolymer or its analogs are added as template D N A ' s with the E. coli polymerase.
Little is known about the de novo synthesis of these polymers. Efforts to detect low-molecular-weight intermediates in the lag period of d A T syn
thesis were not successful.1 0 7 Before any conversion of labeled triphosphates to an acid-insoluble form could be found, trace amounts of macromolecular dAT were detected, early in the lag period, by using aliquots from the reaction mixture to reduce the lag period in a second de novo synthesis.
Evidence t h a t this "lag-reducing activity" was of macromolecular size came from the rate at which the activity sedimented in a preparative ultra- centrifuge. Recently it has been found t h a t the octanucleotide ρ (AT) 4 , synthesized chemically, will reduce the lag period of a de novo synthesis.1 0 8 I t would be interesting to know why the synthetic polymer containing the AT base pair is an alternating copolymer while the GC base pair is
346 ROBERT L. BALDWIN
built into a pair of homopolymer strands. A naturally occurring D N A con
taining chiefly alternating A T sequences has been found by Sueoka1 0 9 (cf.
Swartz et al.10A) in crab testes.
The synthetic D N A polymers provide a strenuous test of the copying accuracy of D N A polymerase, since d A T and d G d C each contain only a single type of base pair. For example, if any G is incorporated during the synthesis of d A T one could detect it with great sensitivity by using radio
active d G T P and unlabeled d A T P and d T T P . I n such an experiment no incorporation of P3 2-labeled d G T P into d A T was found by T r a u t n e r et alno; the sensitivity of detection could be extended to 1 part in 500,000. However when dABU, an analog of d A T containing 5-bromouracil in place of T, was used as a template for further d A B U synthesis, an incorporation of G was detected at a level of one residue of G for every few thousand of A and BU. This fits the hypothesis t h a t the mutagenic action of B U results from an occasional pairing of G with the enol tautomer of B U .1 7 , 1 1 1 However, analysis of the dinucleotide frequencies showed a considerable fraction of the G incorporated next to G. On the hypothesis just given, all the G should have been next to BU. Also, the incorporation of G found when other triphosphates were omitted suggests some end addition.
4. BA S E PAIRING AS THE MECHANISM OF COPYING
On the basis of their structure for D N A Watson and C r i c k1-1 7 suggested t h a t copying of a template D N A could proceed by pairing A with Τ and G with C (Fig. 4). Studies with the purified D N A polymerase from E. coli confirm this and also indicate t h a t the rate of D N A synthesis is negligible unless this base pairing can take place. (Thus, there is no initial synthesis in the absence of a template D N A nor incorporation of G or C when d A T is used as the template.) The first point to be made is t h a t the copying mecha
nism will function without preexisting base pairs in the template since single-stranded D N A ' s will serve as templates. This is true not only of the natural D N A from phage φΧ-174 but also of the synthetic D N A homo- polymers such as d C .1 0 6 Thus, any mechanism is unlikely which uses the entire A T or GC base pair for copying, because then one would need a separate copying mechanism when single-stranded D N A ' s are used as templates.
Second, certain analogs of the natural substrates can be used in the enzymic synthesis.1 1 2 * These include the deoxyribonucleoside triphosphates
* W h e n t r i p h o s p h a t e s c o n t a i n i n g b a s e a n a l o g s are t e s t e d w i t h t h e D N A p o l y m e r a s e s from B. subtilis or T 2 - i n f e c t e d E. coli, t h e s a m e p a t t e r n of r e s u l t s is f o u n d as w i t h t h e E. coli D N A p o l y m e r a s e ,9 3 s u g g e s t i n g t h a t it is a general p a t t e r n for D N A s y n t h e s i s .
7. MECHANISMS OF G E N E REPLICATION 347 of uracil, 5-bromouracil, 5-bromocytosine, and hypoxanthine. I n each case the analog replaces only one of the natural bases, the one predicted from considerations of forming a hydrogen-bonded base pair with the same di
mensions as the A T and GC pairs, and with equivalent positions for the gly
coside
bonds. With one exception the substituted group appears on the outside of the helix. The exception is the replacement of G by hypoxanthine, which is a substitution of hydrogen for the 2-amino group of G and leaves intact the dimensions and two of the three hydrogen bonds of the GC base pair.A third piece of evidence comes from a study of the limited reaction11* which occurs when a template D N A is incubated with D N A polymerase in the absence of one of the triphosphates. I t is found t h a t one or a few nucleotides are added covalently to the 3'-OH end of each primer strand.
When the limited reaction is studied using d A T as the template, the addi
tion only of A or Τ can be detected, not of G or C .1 0 5 This indicates t h a t the limited reaction, like the usual synthetic reaction, proceeds only by specific base pairing.
Base pairing of the template with the newly synthesized D N A can be demonstrated in another way, from melting curves of hybrid D N A ' s con
taining complementary strands of d A T and dABU. Hybrid molecules can be made from a solution containing d A T and d A B U by an annealing proce
dure.1 1 4 I n low salt the A T and A B U base pairs have markedly different thermal stabilities, so t h a t one can follow the melting of d A T : d A B U hybrid molecules in the presence of d A T : d A T and d A B U "dABU, since the hybrid molecules melt in an intermediate temperature zone (Fig. 9b).
Then, when d A T is used as a template for the enzymic synthesis of d A B U or vice versa, it is found t h a t the first product of synthesis melts in the hybrid melting zone (Fig. 9 a ) .1 1 5
The energy of hydrogen bonding between bases is only a few kilocalories per mole of base p a i r s .3 0 , 1 1 6 W h a t then accounts for the extraordinary accuracy of the D N A polymerase in placing A opposite Τ and G opposite C? One possibility is t h a t steric factors are decisive: t h a t a nucleoside triphosphate is accepted if it can fit into an active site which includes both the enzyme and the template D N A , so t h a t the resulting base pair has the correct dimensions and positions for the glycosidic bonds. It is interesting t h a t the D N A polymerase from E. coli will incorporate ribonucleoside triphosphates into a polymer in the presence of Μη"1"4 -.1 1 7 Lee-Huang and Cavalieri1 1 8 have reported t h a t the R N A homopolymer pair rArU, which forms a double helix similar to the D N A helix3 2 b u t whose diameter m a y be somewhat greater,1 1 9 will serve as a template for D N A synthesis using the D N A polymerase from E. coli, and resulting in the synthesis of the D N A homopolymer pair dAdT.
348 ROBERT L. BALDWIN
.301 I I 1 I 1 1 1 1 I I I 1 1 I 1 1 I I I I I I I 22 24 26 26 30 32 34 36 38 40 42. 44
Temperature (°C)
F I G . 9. M e l t i n g c u r v e s of a b s o r b a n c e v s . t e m p e r a t u r e for h y b r i d m o l e c u l e s c o n t a i n i n g d A T a n d d A B U , s h o w i n g t h a t m e l t i n g c u r v e s can be used t o d e m o n s t r a t e d A T : d A B U b a s e pairs a n d t h a t t h e p h y s i c a l l y a n d e n z y m i c a l l y f o r m e d h y b r i d m o l e cules m e l t alike. I n t h e b o t t o m figure t h e h y b r i d m o l e c u l e s were m a d e p h y s i c a l l y b y a n a n n e a l i n g p r o c e d u r e .1 1 4 I n t h e t o p figure t h e y were m a d e e n z y m i c a l l y b y u s i n g d A B U as a t e m p l a t e for d A T s y n t h e s i s .1 1 5 I n b o t h cases m o s t of t h e d A T p r e s e n t m e l t s in a h y b r i d m e l t i n g z o n e 3° C . a b o v e t h e Tm for d A T . T h e d A B U w h i c h is re
l e a s e d on m e l t i n g t h e h y b r i d forms a d A B U : d A B U helix and t h e n m e l t s later w h e n t h e d A B U m e l t i n g zone is r e a c h e d . ( T h e s e c o n d m e l t i n g c u r v e s s h o w t h a t t h e d A T : d A B U b a s e pairs d o n o t re-form after m e l t i n g a n d c o o l i n g in l o w s a l t . T h e 27 w' s differ in t h e t w o figures b e c a u s e of small differences in t h e N a+ c o n c e n t r a t i o n , b u t t h e r e l a t i v e TM}8 for t h e t h r e e s p e c i e s are t h e s a m e in b o t h e x p e r i m e n t s . )
B . SEMICONSERVATIVE REPLICATION
I n addition to suggesting base pairing as the mechanism for copying D N A , the Watson-Crick model4 focused attention on the integrity of the D N A molecule. The question is whether the original helix remains intact after copying (conservative replication) or whether the strands separate during replication, each strand remaining intact (semiconservative replica-
7. MECHANISMS OF GENE REPLICATION 349 tion), or whether the strands themselves are fragmented and rejoined during replication (dispersive replication). These terms were used b y Delbruck and Stent,4 4 who discuss some of their implications. Actually, examples of all three modes of copying D N A now are known: in the DNA-directed synthesis of R N A the D N A helix remains intact after copying (see Section Ι Ι Ι , Ε ) , while the replication of D N A itself appears to be semiconservative but often accompanied by fragmentation and rejoining of chains, especially in the case of the T-even phages.
1. TH E MESELSON-STAHL EXPERIMENT
Meselson and S t a h l3 9 realized t h a t if they could label the newly syn
thesized D N A chains with a lighter isotope, and then separate the D N A species of different densities, they could determine whether the parental D N A remains intact after replication. T h e method of centrifugation to equilibrium in a density gradient,7 2 which was developed for this purpose, has since become one of the major tools of the nucleic acid chemist. I n their experiment E. coli Β was first grown on a synthetic medium containing N1 5H4C 1 . After changing the medium to one containing N1 4 and allowing growth for various times, they lysed the cells with sodium dodecyl sulfate and examined the D N A in a CsCl density gradient (Fig. 10). For molecules of high molecular weight the technique is sufficiently sensitive to resolve components differing in density by less t h a n 0.014 g./ml., which is the difference between N1 4- and N1 5-labeled E. coli D N A .
After one generation all the D N A had a hybrid density. After two genera
tions half the D N A was hybrid, half light (Fig. 10). The results were exactly those predicted for semiconservative replication: Meselson and Stahl had shown t h a t the D N A contained two equal subunits which separate on replication, and t h a t each subunit remained intact through m a n y genera
tions. Further, they showed t h a t the two subunits could be separated b y heating at 100°C. At t h a t time this was a surprising result: it was t h e n widely believed t h a t the two strands of a D N A helix are not disengaged after melting because they remain wound around each other. This belief was based on measurements by light scattering of the molecular weight before and after heating.3 8 Since then, it has been found t h a t there are serious technical problems in measuring the molecular weights of large D N A ' s , especially b y light s c a t t e r i n g .5 1 , 1 2 0·1 2 1
2. STRUCTURE OF THE HY B R I D D N A
T h e Meselson-Stahl experiment gave results in complete agreement with the semiconservative replication of D N A , b u t their results could also fit a special type of conservative model for replication. Measurements of molecu
lar weight and also of the kinetics of enzymic breakdown of D N A led
350 R O B E R T L. BALDWIN
Cavalieri et al}22 to suggest such a model. I n their view each subunit of the Meselson-Stahl experiment is itself a double helix, the two double helices being held together by unspecified bonds which are broken when the D N A is copied at the next replication (for a summary of this and later work, see
ι
F I G . 10. T h e M e s e l s o n - S t a h l e x p e r i m e n t3 9: t h e s e d e n s i t y g r a d i e n t p a t t e r n s s h o w t h e r e l a t i v e a m o u n t s of h e a v y , h y b r i d , a n d l i g h t D N A at v a r i o u s g e n e r a t i o n t i m e s after transferring E. coli g r o w n in an N1 5 m e d i u m t o an N1 4 m e d i u m . T h e e x i s t e n c e of a h y b r i d D N A , i t s p e r s i s t e n c e d u r i n g m a n y g e n e r a t i o n s , a n d t h e r e l a t i v e a m o u n t s of t h e t h r e e s p e c i e s at different g e n e r a t i o n t i m e s s h o w t h a t E. coli D N A c o n t a i n s t w o equal s u b u n i t s w h i c h s e p a r a t e on r e p l i c a t i o n , e a c h s u b u n i t r e m a i n i n g i n t a c t .
Cavalieri and Rosenberg1 2 3). Several studies have been made to clarify this point, most of them concerned with the structure of the hybrid D N A and the nature of the bonds linking the subunits. If the two subunits of the hybrid D N A are the two strands of a D N A helix, then the Meselson-Stahl experiment establishes the semiconservative replication of D N A .
Rolfe1 2 4 used sonic breakage to show t h a t the subunits are not linked
7. MECHANISMS OF GENE REPLICATION 351 end to end. This treatment is believed to break across both strands of a D N A helix, and should yield some heavy and also some light D N A if the hybrid D N A is an end-to-end dimer. Schildkraut et αΖ.52 found t h a t the two subunits separate just at the top of the thermal melting curve of absorbance vs. temperature. The same conclusion was reached by Frei- felder and Davison,1 2 5 who fixed the degree of melting by reaction with formaldehyde and then studied the separation of subunits as a function of the degree of melting. Thus the conditions needed for separation of the hybrid subunits coincide with complete melting of the D N A helix, and with breaking t h e hydrogen bonds between base pairs. Moreover, t h e two subunits cannot melt independently of each other.1 2 6 This was shown with hybrid D N A from E. coli labeled with B U in one subunit. D N A completely labeled with B U melts at a lower p H than unlabeled D N A (cf. Fig. 11a).
Both subunits of the hybrid D N A were found to melt together, at an inter
mediate p H (Fig. l i b ) .
Also, the recombination experiments of M a r m u r , D o t y and their co
workers (Section II,B) are most simply interpreted by assuming t h a t the subunits are the individual strands. After heating the D N A at a tempera
ture where the subunits are separated, it is found t h a t annealing conditions which give specific recombination of the subunits5 2 also produce renatured D N A in which the original helical structure has been largely restored.5 3
Different methods of measuring the number of strands in D N A from actively dividing organisms have given conflicting results. Although electron microscope studies led Hall and Cavalieri1 2 7 to conclude t h a t much of the D N A is four-stranded, measurements of the mass per unit length by low- angle X-ray scattering1 2 8 gave the value for a two-stranded helix on D N A samples prepared from resting and from actively dividing cells, including one sample prepared by Cavalieri.
Cairns' studies of D N A replication in E. c o / i8 3 , 8 4 based on autoradiogra
phy with tritium labeling, add to the evidence t h a t replication is semi- conservative (see Section I I I , C ) . Replication of a more complex type of chromosome, t h a t of the bean plant, was found to be semiconservative by Taylor et al.,129 who used autoradiography. Sueoka1 3 0 has shown t h a t D N A replication in the alga Chlamydomonas follows the p a t t e r n seen in the Meselson-Stahl experiment.
3. In Vitro SYNTHESIS
I t is of considerable interest to know whether D N A synthesis in vitro, using the purified D N A polymerase, will show semiconservative replica
tion. When the newly synthesized D N A is labeled in such a way t h a t it
I 1 I 1 I 1 1 1 I I _ 0 Original pH c h a n q e _ 0 2 n d pH c h a n q e
1 1 I I ι ι ι ι
0 Θ
^.o—o—
ο
1
1 1 1
| 0 - 0* o -•
ό
^ 0 O 0 -fee
dABU I _
- t 1
:
o dABU+dAT^1 1
0
-
I ! 1 1 1 1 1 1 1 1 0^ _ : ι ι
1 I 1 ! 1 I I 1 1 1 I 1 1 ι w: ι , i i l l 7 4 75 &2 6.6 9-0 9.4 9-S 102 10.6 11.0 11.4
( F I G . 11a)
pH Melting oP H y b r i d DNA LH
Hybrid pH P. I
Λ
HI \ / \J Hybrid pH 11,28
i »
1 Λ Denatured Light M J V £ Heavy pH 11.50 LL
Ι Λ Η
/ \ Δ Hybrid £ Native I / V / \J Light pH 11.28
( F I G . l i b )
3 5 2