Genes 2019, 10, genes-581348; doi: FOR PEER REVIEW www.mdpi.com/journal/genes
Article
1
LIP1 regulates the plant circadian clock via the
2
oscillator component GIGANTEA
3
Anita Hajdu1, †, Kata Terecskei1, †, Péter Gyula2, Éva Ádám1, Anna Nyakó1, Orsolya Dobos1, 3, and
4
László Kozma-Bognár 1, 4,*
5
1 Institute of Plant Biology, Biological Research Centre, Szeged H-6726, Hungary; hajdu.anita@brc.mta.hu
6
(A.H.); kata@gmail.com (K.T.); adam.eva@brc.mta.hu (É.Á.); nyako.anna@gmail.com (A.N.);
7
dobos.orsoly@brc.mta.hu (O.D.);e-mail@e-mail.com
8
2 Agricultural Biotechnology Institute, Epigenetics Group, National Agricultural Research and Innovation
9
Center, Gödöllő H-2100, Hungary; gyula.peter@abc.naik.hu (P.G.)
10
3 Doctoral School in Biology, Faculty of Science and Informatics, University of Szeged, Szeged H-6726,
11
Hungary
12
4 Department of Genetics, Faculty of Sciences and Informatics, University of Szeged, Szeged H-6726, Hungary;
13
* Correspondence: kozma_bognar.laszlo@brc.mta.hu (L.K.-B.)
14
† These authors contributed equally to this work
15
Received: June 18, 2019; Accepted: September 19, 2019; Published:
16
Abstract: Circadian clocks are biochemical timers regulating many physiological and molecular
17
processes according to the day/night cycles. The function of the oscillator relies on negative
18
transcriptional/translational feedback loops operated by the so-called clock genes and the encoded
19
clock proteins. The small GTPase LIGHT INSENSITIVE PERIOD 1 (LIP1) is a circadian clock-
20
associated protein that regulates light input to the clock in the model plant Arabidopsis thaliana. In
21
the absence of LIP1, the effect of light on free-running period length is much reduced. We showed
22
that LIP1 is also required for suppressing red and blue light-mediated photomorphogenesis, light-
23
controlled inhibition of endoreplication and tolerance to salt stress. Here we demonstrate that LIP1
24
is present in a complex of clock proteins GIGANTEA (GI), ZEITLUPE (ZTL) and TIMING OF CAB
25
1 (TOC1). LIP1 participates in this complex via GUANINE EXCHANGE FACTOR 7. Analysis of
26
genetic interactions proved that LIP1 affect the oscillator via modulating GI function. Moreover, we
27
showed that GI also connects LIP1 to the regulation of salt stress and endoreplication, but these two
28
proteins control photomorphogenesis by separate routes. Collectively, our results suggest that LIP1
29
attenuates selected functions of GI, possibly by interfering with binding of GI to downstream
30
signalling components.
31
Keywords: Arabidopsis; circadian clock; GIGANTEA, small GTPase LIP1.
32 33
1. Introduction
34
Circadian clocks are endogenous timekeepers that coordinate internal physiological responses to the
35
predicted external environment. In Arabidopsis, 30% of the transcriptome is circadian regulated and
36
this includes processes such as metabolism, the induction of flowering, growth, responsiveness to
37
hormones and biotic and abiotic stress (1-3). Consequently, having an internal oscillator that closely
38
matches external time enhances plant fitness (4). Studies that investigated responsiveness to periodic
39
stress cues or metabolism have highlighted the importance of metabolic oscillations in the regulation
40
of circadian rhythms (5, 6). It has been proposed that endogenous timekeepers not only predict
41
external stress cues, but also provide the basis for temporal segregation between incompatible
42
cellular metabolic processes that would otherwise be energetically futile and stressful (7, 8).
43
Plant circadian rhythms are generated through a series of transcriptional-translational loops. At the
44
center of the oscillator is a repressive feedback loop formed between the morning expressed MYB
45
transcription factors CIRCADIAN CLOCK ASSOCIATED 1 (CCA1) and LATE ELONGATED
46
HYPOCOTYL (LHY) and the night-phased TIMING OF CAB EXPRESSION 1 (TOC1), also known as
47
PSEUDO RESPONSE REGULATOR 1 (PRR1) (9, 10). This core loop is subsequently regulated by a
48
series of morning and evening loops. In the morning, sequential expression of PRR9/7/5 starting with
49
PRR9 just after dawn results in the repression of CCA1/LHY expression throughout the day and early
50
evening (11). At dusk, GIGANTEA (GI) and ZEITLUPPE (ZTL) co-associate to degrade TOC1 (12,
51
13), and the evening complex(14), composed of EARLY FLOWERING3, EARLY FLOWERING4 and
52
LUX ARRYTHMO (LUX), represses the expression of GI, LUX, PRR9 and PRR7 (15, 16).
53
The small GTPase LIGHT INSENSITIVE PERIOD 1 (LIP1) is a circadian clock- associated protein
54
that regulates light input to the clock in the model plant Arabidopsis thaliana. In the absence of LIP1,
55
the effect of light on free-running period length is much reduced. We showed that LIP1 is also
56
required for suppressing red and blue light-mediated photomorphogenesis, light-controlled
57
inhibition of endoreplication and tolerance to salt stress. Here we demonstrate that LIP1 is present
58
in a complex of clock proteins GIGANTEA (GI), ZEITLUPE (ZTL) and TIMING OF CAB 1 (TOC1).
59
LIP1 participates in this complex via GUANINE EXCHANGE FACTOR 7. Analysis of genetic
60
interactions proved that LIP1 affect the oscillator via modulating GI function. Moreover, we showed
61
that GI also connects LIP1 to the regulation of salt stress and endoreplication, but these two proteins
62
control photomorphogenesis by separate routes. Collectively, our results suggest that LIP1
63
attenuates selected functions of GI, possibly by interfering with binding of GI to downstream
64
signalling components.
65
2. Materials and Methods
66
2.1. Plant materials and growth conditions.
67
The Arabidopsis (Arabidopsis thaliana) ecotype Columbia-0 (Col-0) was used as the background
68
for all the experimental lines. pifQ (pif1pif3pif4pif5), pifQ CCA1:LUCIFERASE (LUC) and wild-type
69
(wt) CCA1:LUC transgenic plants are described in (Leivar et al., 2008; Shor et al. 2017). The PIF-
70
overexpression (PIF-ox) lines used were 35spro:PIF1-HA (Zhu et al., 2015), 35spro:PIF3-myc (Park et
71
al., 2004), 35spro:PIF4-myc and 35spro:PIF5-myc (Sakuraba et al., 2014). For all experiments, seeds
72
were imbibed and cold treated at 4°C for 4 days and sown onto Petri dishes with Murashige and
73
Skoog (MS) medium (Duchefa Biochemie, Netherlands) with or without 3% (w/v) sucrose (for
74
luciferase assay), or 2% sucrose (w/v) (for leaf movement assays and RT-PCR). Unless otherwise
75
stated, plants were grown in 14 hours light: 10 hours dark (14 L:10 D) with 100 μmol m−2 s−1 white
76
light supplied by Philips fluorescent lights TLD 18W/840 at 23°C.
77
2.2. Bioluminescence assays.
78
For the circadian bioluminescence assays, 6-8 seedlings from each of 3-4 independent lines
79
carrying the CCA1:LUCIFERASE (CCA1:LUC) reporter (pifQ CCA1:LUC and wt CCA1:LUC) were
80
grown for 8 days in 14 L:10 D. After spraying with 2.5mM luciferin (D-Luciferin, Potassium salt, Gold
81
Biotechnology, St Louis, MO, USA) in 0.01% Triton X-100 the seedlings were transferred to a growth
82
chamber mounted with a Hamamatsu ORCA II ER CCD camera (C4742-98 ERG; Hamamatsu
83
Photonics, Hamamatsu City, Japan). Light was provided by red and blue light emitting diodes
84
(LEDs), with regulated total fluence rates. Luciferase activity was imaged every two hours for at least
85
four days. Images were analyzed with ImagePro software (Media Cybernetics, Inc., Bethesda, MD,
86
USA). Data were imported into the Biological Rhythms Analysis Software System (BRASS;
87
http://www.amillar.org) and analyzed with the FFT-NLLS (Fourier Transform-NonLinear Least
88
Squares) suite of the program, as previously described (Plautz et al., 1997). Rhythms with a period
89
between 15 and 35 hours were taken to be within the circadian range.
90
2.3. RNA isolation and quantitative RT-PCR.
91
Ten to twelve seedlings were harvested per sample and total RNA extracted as previously
92
described (Green and Tobin, 1999). RNA samples were treated with DNase I (PerfeCTa DNAse from
93
Quanta bio, Beverly, MA, USA) according to the manufacturer’s instructions. From each DNA-free
94
RNA sample, 5 μl aliquots were used as a template to produce cDNA, using the qScript cDNA
95
SuperMix (Quanta bio). 2.5 μl of template cDNA was used for quantitative RT-PCR reaction with
96
SYBR green reagent (KAPA SYBR FAST qPCR kit Master Mix, Kapa Biosystems, MA, USA) according
97
to the supplier’s protocol. Three technical repeats were made for each sample. Fluorescence was
98
detected using the QuantStudio 12K Flex system (Thermo Fisher Scientific, MA, USA). PROTEIN
99
PHOSPHATASE 2A (PP2A, AT1G13320), was used as a control for normalization (Czechowski et al.,
100
2005). Quantitation calculations were carried out using the 2–ΔΔCT formula as described (Nozue et
101
al., 2007). The primers are shown in Supplementary Table 1.
102
2.4. Yeast two-hybrid tests for protein interactions.
103
Yeast cells were co-transformed with different cDNA-fragments cloned into pGADT7 and
104
pGBKT7 vectors (Clontech) in frame with the GAL4 transcriptional activator domain or with the GAL4
105
DNA-binding domain, respectively. Transformed cells were plated on Leu-/Trp- (LW) and Ade-/Leu-
106
/Trp- (ALW) CSM agar plates and were grown at 30 oC for 5 days. For -galactosidase enzyme activity
107
assay, three independent colonies were picked up from LW or ALW plates and inoculated into LW
108
CSM liquid media, and were shaken at 30 oC until density reached OD600 0.8, when the assay (using
109
O-nitrophenyl-β-D-galactopyranoside as substrate) was carried out (Yasuhara et al, 2004). For
110
documentation of the growth on selective medium, single colonies from LW or ALW plates were
111
diluted in 100 l of sterile water and 5 ls of them were dropped on fresh LW and ALW plates. After 5
112
days of growth plates were scanned by a flat-bed scanner. Expression of the mutated ZTL proteins in
113
yeast were confirmed by Western blotting using anti c-Myc and anti HA antibodies (Sigma).
114
3. Results
115
3.1.1. Pattern and level of clock gene expression in the lip1-2 mutant
116
It has been demonstrated previously that LIP1 affects the circadian clock by mediating light
117
signalling to the oscillator (Kevei 2007). Input light signals may affect the level, the activity or
118
subcellular localization of oscillator components to set the pace and phase of the circadian clock. In
119
order to test the effect of LIP1 on clock gene expression, Col wild type and lip1-2 mutant seedlings
120
were entrained to 12 h light:12 h dark (12:12 LD) photocycles for a week and then transferred to
121
continuous red light at relatively low fluence rate (5 μmol m-2 s-1), where the short period phenotype
122
of lip1 mutants is readily detectable (Kevei 2007)(Terecskei 2013). The accumulation pattern and level
123
of selected clock genes was analysed by qPCR assays. The genes were selected to represent the
124
different regulatory loops, but also to include the first identified main components (CCA1, TOC1),
125
genes with sequential peak times during the day (PRR5, 7, 9) and key elements of the Evening
126
Complex (LUX, ELF4) as well. Figure 1 shows that rhythmic accumulation of all tested genes
127
displayed shorter periods in the mutant compared with the wild type control. However, mRNA
128
levels did not change consistently in the mutant. These data indicate that altered level of clock gene
129
expression probably does not underlay the period phenotype of lip1-2, thus the primary and direct
130
effect of LIP1 on the oscillator is not the transcriptional regulation of clock genes.
131
3.1.2. Identification of proteins interacting with LIP1
132
Since the results above suggested that LIP1 may exerts its clock-related function at
133
posttranscriptional/protein level, we aimed at identifying proteins thorough which this regulation
134
could take place. First we tested the interactions between LIP1 and several clock proteins (CCA1,
135
TOC1, GI, ZTL, ELF4, ELF3, LUX, PRR9) using the GAL4-based yeast two-hybrid (Y2H) system
136
without any positive results (data not shown). To expand the range of potential partners, in the next
137
step we performed a yeast two-hybrid (Y2H) screen employing LIP1 as bait. We isolated 7 clones
138
encoding protein fragments that interacted with LIP1 in a reproducible manner. However, only one
139
of these retained the ability for interaction when the corresponding full-length protein was co-
140
expressed with LIP1 (Figure 2A). The gene is designated as AT5G02010 and encodes for ROP (RHO
141
OF PLANTS) GUANINE NUCLEOTIDE EXCHANGE FACTOR 7 (ROPGEF7, GEF7 hereafter in the
142
text). GEF7 belongs to the family of ROPGEF proteins consisting of 14 members in Arabidopsis. These
143
proteins facilitate the replacement of GDP by GTP bound to the plant-specific Rop GTPases, leading
144
to the activation of these signalling factors. The LIP1-GEF7 interaction was verified by a Luciferase
145
Complementation Assay (Figure 2B) in E. coli cells, overcoming the problem of transactivation by
146
GEF7 in the Y2H system. This result also suggested that no plant-specific posttranslational
147
modifications are required for establishment of LIP1-GEF7 interaction. To reveal potential links to
148
the oscillator, we tested interactions between GEF7 and clock proteins CCA1, TOC1, GI, ZTL, ELF4,
149
ELF3, LUX and PRR9. Significant binding to LIP1 was detected in the case of GI, TOC1 and ZTL
150
(Fig.2C-E). Interestingly, it has been demonstrated earlier that ZTL plays a role in the degradation of
151
TOC1 (Mas 2003), whereas GI stabilizes ZTL in a light dependent manner (Kim 2007) via direct
152
interactions. We verified these interactions in the Y2H system (Fig.2G, H), but also demonstrated
153
physical association between TOC1 and GI (Fig.2F) that has not been reported before. These data
154
indicate that despite the lack of direct interactions with oscillator components, LIP1 may be present
155
in clock protein complexes through GEF7.
156 157
3.1.3. Genetic analysis identifies GI as the clock component targeted by LIP1
158
In order to reveal the functional consequences of the indirect interactions described above and
159
to test if one of the complex-forming clock proteins represents the entry point of LIP1-derived in the
160
oscillator, double mutants were generated by crossing lip1-2 to gi-101, toc1-4, ztl-3 or cca1-1. The
161
cca1-1 mutant was used as control, since neither direct nor indirect interaction was detected between
162
CCA1 and LIP1. The mutant combinations carried the CCR2:LUC or the CAB2:LUC reporters
163
facilitating the analysis of the circadian phenotypes. Plants, including the wild type and the single
164
mutant controls were assayed in low intensity red light (Fig.3). Visual inspection of rhythmic traces
165
indicated additive period phenotypes for lip1-2 and toc1-4 (Fig.3B), ztl-3 (Fig.3C) and cca1-1 (Fig.3D).
166
Estimates of free-running periods verified this observation with quantitative data (Table 1). The
167
period of lip1-2 ztl-3 was in between the two parent singles, whereas periods of lip1-2 toc1-4 and lip1-
168
2 cca1-1 were significantly shorter compared with the parental lines. In contrast, lip1-2 gi-101
169
produced CCR2:LUC rhythms with periods indistinguishable from that of gi-101, but significantly
170
longer than that of the lip1-2 single (Fig.3A, Table 1). Moreover, the reduction of amplitude seen in
171
gi-101 was also clearly observable in lip1-2 gi-101. These data demonstrate that GI is epistatic to LIP1
172
in the regulation of the circadian oscillator.
173
3.1.4. Functions of LIP1 in the regulation of endoreplication and salt stress responses are mediated via GI
174
In addition to its function in the regulation of the circadian clock, LIP1 was shown to control
175
ploidy levels, responses to salt stress and light-dependent hypocotyl elongation (Terecskei 2013).
176
Since we showed that LIP1 affects the clock through GI, we aimed at testing if other phenotypes of
177
the lip1-2 mutants also depend on GI.
178
Previously we demonstrated that the rounded shape of epidermal pavement cells of lip1 mutants is
179
due to increased ploidy levels, which is the result of impaired suppression of endoreplication
180
(Terecskei 2013). We monitored this phenotype as a proxy for ploidy levels in the different genetic
181
backgrounds. Figure 4 illustrates that in agreement with previous results the shape of pavement cells
182
in 7-day-old light-grown lip1-2 seedlings (Fig.4B) was dramatically different from that of the WT
183
plants (Fig.4A). The gi-101 mutant did not show obvious alterations compared with the WT (Fig.4C).
184
Interestingly, the lip1-2 gí-101 double mutant (Fig.4D) phenocopied the gi-101 single indicating the
185
functional GI is required for the expression of the ploidy phenotype of lip1-2.
186
LIP1 was shown be required for efficient tolerance of high salinity as germination and development
187
of lip1 mutants was severely impaired at 100 mM NaCl, which was clearly tolerated by WT plants
188
(Figure 5) (Terecskei 2013). The gi-101 mutant did not show any symptoms in these conditions, in
189
agreement with previous reports (Kim 2013)(Sakuraba 2017). The lip1-2 gi-101 double mutant
190
behaved like gi-101 indicating that GI functions downstream of LIP1 in controlling salt stress
191
responses.
192
3.1.5. Functions of GI in the regulation of photomorphogenesis and flowering time are not affected by LIP1
193
Both LIP1 and GI play a role in light-controlled hypocotyl elongation, although they exert
194
opposite effects: compared with WT, lip1 or gi mutants produce shorter or longer hypocotyls,
195
respectively, when grown in continuous red or blue light (Martin-Tryon 2007)(Kevei 2007)(Terecskei
196
2013). However, none of them are involved in far-red light signal transduction (Huq 2000) (Terecskei
197
2013). To test genetic interaction between LIP1 and GI for these phenotypes, hypocotyl lengths of
198
seedlings of different genotypes were determined after 4 days of growth in continuous red, blue and
199
far-red light and in darkness. In order to reflect the light-controlled component of hypocotyl
200
elongation, height of light-grown seedlings was normalized to the height of the corresponding dark-
201
grown plants. Figure 6A shows that regarding the inhibition of hypocotyl elongation by red and blue
202
light lip1-2 was hypersensitive, whereas gi-101 was hyposensitive, as reported earlier. The lip1-2 gi-
203
101 double produced hypocotyl lengths intermediate between the two single mutants and mimicking
204
WT. As expected, none of the mutants showed alterations from the WT in far-red light.
205
GI is a key player of photoperiodic flowering upregulating FT transcription by CO-dependent (Sawa
206
2007) and CO-independent (Sawa 2011) routes. Accordingly, flowering is dramatically delayed in gi
207
mutants. Owing to the lack of information on the flowering phenotype of lip1 mutants, plants of the
208
different genotypes were grown in short day (8 h light/16 h dark, SD) or long day (16 h light/8 h dark,
209
LD) conditions. The time of flowering was determined as the number of rosette leaves at bolting.
210
Figure 6B demonstrates that flowering time was not altered in the lip1-2 mutant in neither conditions.
211
The gi-101 single flowered later then the WT, whereas the lip1-2 gi-101 double was indistinguishable
212
from the gi-101 single.
213
These results suggest that GI and LIP1 regulate hypocotyl elongation via largely different signalling
214
routes and that the function of GI in flowering time initiation is not modulated by LIP1.
215
3.2. Figures, Tables and Schemes
216
217
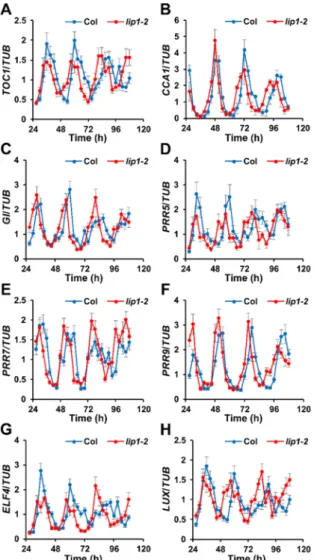
Figure 1. mRNA accumulation pattern of clock genes in the lip1-2 mutant
218
Wild type (Col) and lip1-2 mutant seedlings were grown in 12 h light: 12 h dark photocycles for 7 days and
219
transferred to continuous red light (5 µmol m-2 s-1). Samples were harvested at 3-hour intervalls, starting 27 h
220
after the transfer. mRNA levels of TOC1 (A), CCA1 (B), GI (C), PRR5 (D), PRR7 (E), PRR9 (F), ELF4 (G),
221
and LUX (H) were determined by qPCR and normalised to the corresponding TUB mRNA levels. Average
222
values of 3 indpendent replicates are plotted, error bars represent Standard Error values.
223
224
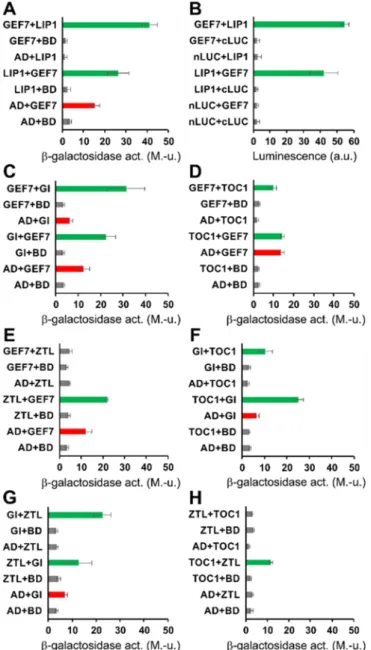
Figure 2. Identification of proteins directly or indirectly interacting with LIP1
225
Full-length LIP1, GEF7, GI, TOC1 and ZTL proteins fused to the transcriptional activation domain (AD) or the
226
DNA-binding domain (BD) of the GAL4 transcription factor were coexpressed in yeast (PJ69-4A) cells (A, C-
227
H). Interactions were tested in either (AD and BD) configurations. For each combination, the first or the second
228
indicated protein carried the AD or the BD fusion tag, respectively. AD and BD correspond to controls, where
229
these GAL4 derivatives were expressed without foreign fusion partners. β-galactosidase activity, reporting the
230
activation of the lacZ marker and therefore the strength of interaction between the two given fusion proteins,
231
was determined from liquid-cultured transformant cells. Green bars indicate activation above the background
232
levels (grey bars). GEF7 and GI fused to BD were able to activate the markergene without any interacting
233
partners (transactivation, red bars). The assays were repeated 3-4 times with essentially the same results. Error
234
bars represent Standard Error values of 3 technical repeats of a representative assay. M.-u.: Miller-units.
235
The interaction between LIP1 and GEF7 was also tested by luciferase complementation assays (B). LIP1 or
236
GEF7 fused to the N- or C-terminal fragment of firefly luciferase (nLUC or cLUC) in either configurations
237
were coexpressed in E. coli BL21 Rosetta cells along with the corresponding controls. Luminesce of bacterial
238
patches (5 patches for each combinations) was detected by a cooled CCD camera. Error bars represent the
239
Standard Error values of 3 independent assays. a.u.: arbitrary units = counts/patch/5 min.
240
241
242
Figure 3. LIP1 requires GI to affect the circadian clock
243
Seedlings of the indicated genotypes carrying CCR2:LUC (A, C, D) or CAB2:LUC (B) reporter genes were
244
grown in 12 h light: 12 h dark photoperiods for 7 days and transferred to continuous red light (5 µmol m-2 s-1),
245
where luminescence was monitored. For each individual seedlings, values were normalised to the average of
246
all counts collected during the course of the assay. The means of normalised data from 24 seedlings for each
247
genotypes are plotted. Experiments were repeated 3 or 4 times.
248 249
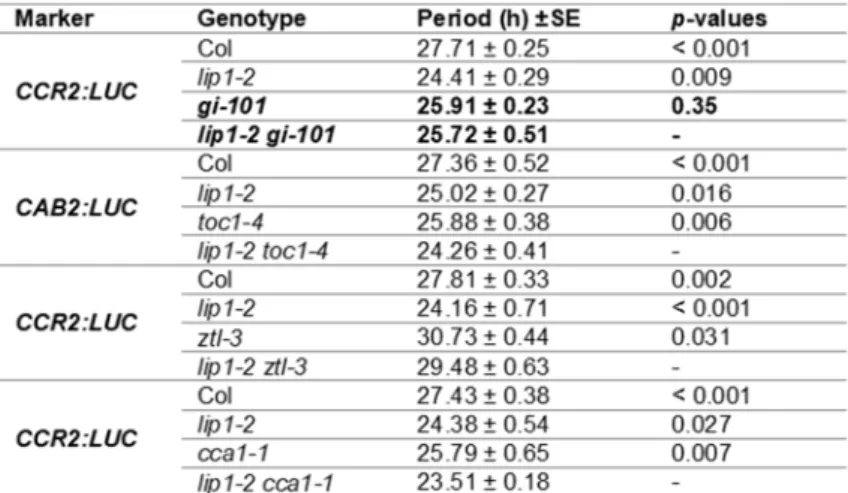
Table 1. Period estimates demonstrate genetic interaction between LIP1 and GI.
250
Luminescence data of plants shown in Figure 3 were analysed by the BRASS3 software package. Free-running
251
periods were estimated by FFT-NLLS analysis. p-values were calculated from pairwise t-tests to determine the
252
significance of differences from the corresponding double mutant in terms of periods.
253
254
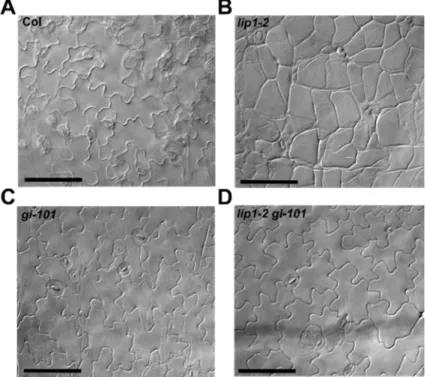
Figure 4. The pavement cell morphology phanotype of lip1-2 mutants is dependent on GI.
255
Pavement cell morphology of Col (A), lip1-2 (B), gi-101 (C) and lip1-2 gi-101 plants grown in 12 h light: 12h
256
dark photocycles for 7 days. Scale bars: 100 µm.
257 258
259
Figure 5. The lack of GI function supresses the salt sensitivity phenotype of lip1-2 mutants.
260
Col, lip1-2, gi-101 and lip1-2 gi-101 seedlings were grown in 12 h light: 12 h dark photocycles for 14 days on
261
media with or without 100mM NaCl.
262
263
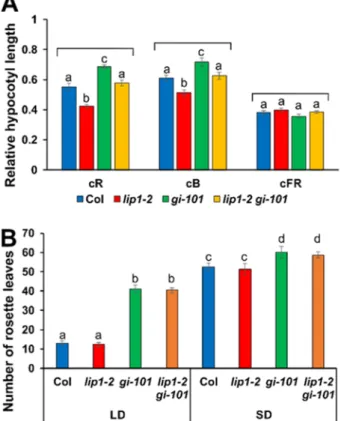
Figure 6. LIP1 affect photomorphogenic responses independently of GI
264
(A) Wild type Col, lip1-2, gi-101 and lip1-2 gi-101 mutant seedlings were grown in continuous red (cR, 20
265
µmol m-2 s-1), blue (cB, 2 µmol m-2 s-1) or far-red (cFR, 1 µmol m-2 s-1) light for 4 days. Hypocotyl lengths were
266
measured and normalised to the corresponding dark-grown hypocotyl lengths. 30-40 seedlings were analysed
267
for each genotype and fluence rate. Error bars indicate Standard Error, and different letters show significant
268
differences at P < 0.05 (Duncan’s test).
269
(B) Plants were grown in 16 h light: 8 h dark (LD) or 8 h light: 16 h dark (SD) photocycles. Rosette leaves were
270
counted when inflorescences reached 1 cm. 12-15 plants were analysed for each genotypes and conditions.
271
Error bars indicate Standard Error, and different letters show significant differences at P < 0.01 (Duncan’s test).
272 273
4. Discussion
274
LIP1 is the first small GTPase that has been functionally linked to the circadian clock in plants.
275
Lack of LIP1 function results in an accelerated circadian oscillator producing short period rhythms.
276
However, the mechanism by which LIP1 affects the oscillator remained unknown. In the present
277
work we aimed at revealing an essential piece of this regulation and identify the particular oscillator
278
component that is primarily targeted by LIP1.
279
We tested the mRNA accumulation of several key clock genes in the loss-of-function allele lip1-
280
2 plants in continuous low fluence red light, where the short period phenotype is most pronounced.
281
No significant changes in mRNA levels of clock genes were found suggesting that transcriptional
282
modulation is not the principal effect of LIP1 on the clock. The pace of the oscillator can also be
283
influenced by altering the function or the turnover of one or more clock proteins. This effect may be
284
mediated via protein-protein interactions. Although direct interaction between LIP1 and clock
285
proteins was not detected, a search for binding partners identified a guanine exchange factor (GEF7),
286
which in turn showed physical interaction with GI, TOC1 and ZTL proteins. GEF7 is a member of the
287
RopGEF protein family and acts as a functional guanine exchange factor to activate Rop GTPase
288
AtRAC1, required for root meristem maintenance (Chen 2011). Interestingly, pairwise interactions
289
between GI, TOC1 and ZTL was also found in Y2H. Supposing that these interactions take place in
290
vivo, the results suggest the existence of a multiprotein complex to which LIP1 may bind through
291
GEF7. It is notable that these factors, including LIP1, operate in the evening. To link LIP1 to the
292
oscillator, one can assume that the complex of GI-TOC1-ZTL could modulate the activity of LIP1
293
through GEF7 and then LIP1 affect the function of one or more clock proteins. Alternatively, LIP1,
294
brought in proximity by GEF7, could influence the activities of GI, TOC1 or ZTL. Analysis of epistasis
295
between LIP1 and the three clock components supported the latter option and identified GI as the
296
downstream target of LIP1-derived signalling to the oscillator. Moreover, we showed that in addition
297
to the circadian phenotype, the ploidy and salt stress phenotypes of lip1 mutants also depend on GI.
298
However, the clear additivity of photomorphogenic phenotypes of gi and lip1 mutants demonstrated
299
that LIP1 does not act exclusively through the modulation of GI function.
300
A simple way to control the function of GI is to regulate the level/turnover or the subcellular
301
localisation of the protein. However, these changes to the GI protein should alter all functions of GI,
302
including flowering time determination (Mizoguchi 2005)(Günl 2009), which phenotype was not
303
observed in lip1 mutants. This indicates that LIP1 does not affect the function of GI in general, but
304
probably acts selectively on particular branches of GI signalling. GI has diverse pleiotropic functions,
305
which appear to be realized via specific protein-protein interactions (see the Introduction). Based on
306
our current results and considering published data we hypothesise that LIP1 affects interaction of GI
307
with downstream effector proteins that relay the effect of GI on the clock and salt stress tolerance.
308
The fact that the cell shape/ploidy phenotype of lip1-2 is supressed in the lip1-2 gi-101 double mutant,
309
strongly indicates that GI plays a role in the regulation of this process as well. This function of GI has
310
not been described and the related specific interactors were not identified yet, but our results suggest
311
that the corresponding hypothetic interactions are also impacted by LIP1. In contrast, interactions
312
representing outputs of GI towards the regulation of photomorphogenesis and flowering should not
313
be affected by LIP1.
314
GI regulates the circadian oscillator via at least two distinct mechanisms. GI triggers
315
destabilization of PIF transcription factors, and thus relieves repression on CCA1 transcription
316
(Nohales 2019). On the other hand, GI stabilizes ZTL in the light, which in turn mediate degradation
317
of clock proteins, among them TOC1, in the dark (Cha 2017). Since CCA1 mRNA levels were not
318
altered in lip1-2 and ztl-3 was not epistatic to lip1-2, we conclude that these two regulatory
319
mechanisms of GI are probably not influenced by LIP1. Rather, these data indicate the existence of
320
an additional, LIP1-modulated functional link from GI to the clock.
321
Plants overexpressing GI (GI-OX) show increased sensitivity to salt stress (Kim 2013) and
322
display circadian rhythms with shortened periods (Mizoguchi 2005). These phenotypes of GI-OX
323
plants are shared with lip1 mutants (Kevei 2007)(Trecskei 2013). Thus we conclude that LIP1
324
attenuates these functions of GI. This very likely holds true of the cell shape phenotype as well, so
325
one could predict that epidermal pavement cells of GI-OX plants have rounded shape, similarly to
326
those of the lip1 mutants.
327
In summary, we demonstrated that LIP1 affects the circadian clock, salt stress responses and
328
epidermal cell shaping via the inhibition of selected functions of GI.
329
Author Contributions: conceptualization, L.K-B. and A.H.; methodology, A.H., K.T., P.G., É.Á., A.N. and O.D.;
330
validation, L.K-B., A.H. and T.K.; formal analysis, P.G., É.Á. and O.D.; writing—original draft preparation, A.H.
331
and T.K.; writing—review and editing, L.K-B.; funding acquisition, L.K-B.
332
Funding: This research was funded by National Research, Development and Innovation Office, grant numbers
333
GINOP-2.3.2-15-2016-00001, GINOP-2.3.2-15-2016-00015 and GINOP-2.3.2-15-2016-00032. The APC was funded
334
by GINOP-2.3.2-15-2016-00001.
335
Conflicts of Interest: The authors declare no conflict of interest.
336 337
References
338
1. Alabadi D, Oyama T, Yanovsky MJ, Harmon FG, Mas P, Kay SA. Reciprocal regulation between TOC1and
339
LHY/CCA1 within the Arabidopsis circadian clock. Science. 2001. 293: 880–883.
340
2. Baudry A, Ito S, Song YH, Strait AA, Kiba T, Lu S, Henriques R, Pruneda-Paz JL, Chua NH, Tobin EM, Kay
341
SA, Imaizumi T. F-box proteins FKF1 and LKP2 act in concert with ZEITLUPE to control Arabidopsis
342
clock progression. Plant Cell. 2010. 22 (3): 606-22.
343
3. Czechowski T, Stitt M, Altmann T, Udvardi MK, Scheible WR. Genome-wide identification and testing of
344
superior reference genes for transcript normalization in Arabidopsis. Plant Physiol. 139, 5–17
345
4. Devlin PF, Kay SA. Cryptochromes are required for phytochrome signaling to the circadian clock but not
346
for rhythmicity. Plant Cell. 2000. 12 (12): 2499-2510.
347
5. Frank A, Matiolli CC, Viana AJC, Hearn TH, Kusakina J, Belbin FE, Newman DW, Yochikawa A, Cano-
348
Ramirez DL, Chembath A, Cragg-Barber K, Hadon MJ, Hotta CT, Vincentz M, Webb AAR, Dodd AN.
349
Circadian entrainment in Arabidopsis by the sugar-responsive transcription factor bZIP63. 2018. Curr Biol.
350
28: 2597-2606.
351
6. Franklin KA, Lee SH, Patel D, Kumar SV, Spartz AK, Gu C, Ye S, Yu P, Breen G, Cohen JD, Wigge PA, Gray
352
WM. Phytochrome-interacting factor 4 (PIF4) regulates auxin biosynthesis at high temperature. Proc Natl
353
Acad Sci U S A. 2011. 108 (50): 20231-5.
354
7. Garner WW, Allard HA. Effect of the relative length of day and night and other factors of the environment
355
on growth and reproduction in plants. J Agric. Res. 1920. 18: 553–606.
356
8. Gould PD, Locke JC, Larue C, Southern MM, Davis SJ, Hanano S, Moyle R, Milich R, Putterill J, Millar AJ,
357
Hall A. The molecular basis of temperature compensation in the Arabidopsis circadian clock. Plant Cell.
358
2006. 18: 1177–1187.
359
9. Gray JA, Shalit-Kaneh A, Chu DN, Hsu PY, Harmer SL. The REVEILLE Clock Genes Inhibit Growth of
360
Juvenile and Adult Plants by Control of Cell Size. Plant Physiol. 2017. 173 (4): 2308-2322.
361
10. Green RM, Tobin EM. Loss of the circadian clock-associated protein 1 in Arabidopsis results in altered
362
clock-regulated gene expression. Proc Natl Acad Sci USA. 1999. 96 (7): 4176-4179.
363
11. Greenham K, McClung CR. Integrating circadian dynamics with physiological processes in plants. Nature
364
Reviews Genetics. 2015. 16 (10): 598-610.
365
12. Haydon MJ, Mielczarek O, Robertson FC, Hubbard KE, Webb AA. Photosynthetic entrainment of the
366
Arabidopsis thaliana circadian clock. Nature. 2013. 502 (7473): 689-692.
367
13. Haydon MJ, Mielczarek O, Frank A, Román Á, Webb AA. Sucrose and ethylene signaling interact to
368
modulate the circadian clock. Plant Physiol. 2017. 175(2):947-958.
369
14. Hsu PY, Harmer SL. Wheels within wheels: the plant circadian system. Trends Plant Sci. 2014. 19 (4): 240-
370 371
9.15. Kim J, Geng R, Gallenstein RA, Somers DE. The F-box protein ZEITLUPE controls stability and
372
nucleocytoplasmic partitioning of GIGANTEA. Development. 2013. 140 (19): 4060-9.
373
16. Kim WY, Fujiwara S, Suh SS, Kim J, Kim Y, Han L, David K, Putterill J, Nam HG, Somers DE. ZEITLUPE
374
is a circadian photoreceptor stabilized by GIGANTEA in blue light. Nature. 2007. 449 (7160): 356-60.
375
17. Knight H, Thomson AJ, McWatters HG. Sensitive to freezing6 integrates cellular and environmental inputs
376
to the plant circadian clock. Plant Physiol. 2008. 148 (1): 293-303.
377
18. Koini MA, Alvey L, Allen T, Tilley CA, Harberd NP, Whitelam GC, Franklin KA. High temperature-
378
mediated adaptations in plant architecture require the bHLH transcription factor PIF4. Curr Biol. 2009. 19
379
(5): 408-13.
380
19. Kumar SV, Lucyshyn D, Jaeger KE, Alós E, Alvey E, Harberd NP, Wigge PA. Transcription factor PIF4
381
controls the thermosensory activation of flowering. Nature. 2012. 484 (7393): 242-245.
382
20. Leivar P, Monte E. PIFs: systems integrators in plant development. Plant Cell. 2014. 26: 56–78.
383
21. Leivar P, Monte E, Oka Y, Liu T, Carle C, Castillon A, Huq E, Quail PH. Multiple phytochrome-interacting
384
bHLH transcription factors repress premature seedling photomorphogenesis in darkness. Curr Biol. 2008.
385
18 (23): 1815-23.
386
22. Lu SX, Knowles SM, Andronis C, Ong MS, Tobin EM. CIRCADIAN CLOCK ASSOCIATED1 and LATE
387
ELONGATED HYPOCOTYL function synergistically in the circadian clock of Arabidopsis. Plant Physiol.
388
2009. 150 (2): 834-43.
389
23. Ma D, Li X, Guo Y, Chu J, Fang S, Yan C, Noel JP, Liu H. Cryptochrome 1 interacts with PIF4 to regulate
390
high temperature-mediated hypocotyl elongation in response to blue light. Proc Natl Acad Sci U S A. 2016.
391
113 (1): 224-9.
392
24. Más P. Circadian clock function in Arabidopsis thaliana: time beyond transcription. Trends Cell Biol. 2008.
393
18 (6): 273-81.
394
25. Nolte C, Staiger D. RNA around the clock - regulation at the RNA level in biological timing. Front. Plant
395
Sci. 2015. 6: 311
396
26. Nozue K, Covington MF, Duek PD, Lorrain S, Fankhauser C, Harmer SL, Maloof JN. Rhythmic growth
397
explained by coincidence between internal and external cues. Nature. 2007. 448 (7151): 358-61.
398
27. Paik, I., Kathare, P.K., Kim, J-I. and Huq, E. (2017) Expanding roles of PIFs in signal integration from
399
multiple processes. Mol Plant 10 (8): 1035-1046.
400
28. Park E, Kim J, Lee Y, Shin J, Oh E, Chung WI, Liu JR, Choi G. Degradation of phytochrome interacting
401
factor 3 in phytochrome-mediated light signaling. Plant and Cell Physiology. 2004. 45: 968–975.
402
29. Pedmale UV, Huang SS, Zander M, Cole BJ, Hetzel J, Ljung K, Reis PA, Sridevi P, Nito K, Nery JR, Ecker
403
JR, Chory J. Cryptochromes Interact Directly with PIFs to Control Plant Growth in Limiting Blue Light.
404
Cell. 2016. 164 (1-2): 233-45.
405
30. Plautz JD, Straume M, Stanewsky R, Jamison CF, Brandes C, Dowse HB, Hall JC, Kay SA. Quantitative
406
analysis of Drosophila period gene transcription in living animals. J Biol Rhythms. 1997. 12 (3): 204-217.
407
31. Romanowski A and Yanovsky MJ. Circadian rhythms and post-transcriptional regulation in higher plants.
408
Front. Plant Sci. 2015. 6: 437.
409
32. Sakuraba Y, Jeong J, Kang MY, Kim J, Paek NC, Choi G. Phytochrome-interacting transcription factors PIF4
410
and PIF5 induce leaf senescence in Arabidopsis. Nature Communications. 2014. 5: 4636.
411
33. Salomé PA, McClung CR. PSEUDO-RESPONSE REGULATOR 7 and 9 are partially redundant genes
412
essential for the temperature responsiveness of the Arabidopsis circadian clock. Plant Cell. 2005. 17 (3): 791-
413
803.
414
34. Sawa M, Nusinow DA, Kay SA, Imaizumi T. FKF1 and GIGANTEA complex formation is required for day-
415
length measurement in Arabidopsis. Science. 2007. 318 (5848): 261-5.
416
35. Smith AM, Zeeman SC, Smith SM. Starch degradation. Annu Rev Plant Biol. 2005. 56: 73-98.
417
36. Shor E, Paik I, Kangisser S, Green R, Huq E. PHYTOCHROME INTERACTING FACTORS mediate
418
metabolic control of the circadian system in Arabidopsis. New Phytol. 2017. doi: 10.1111/nph.14579.
419
37. Somers DE, Devlin PF, Kay SA. Phytochromes and cryptochromes in the entrainment of the Arabidopsis
420
circadian clock. Science. 1998. 282 (5393): 1488-90.
421
38. Sun J, Qi L, Li Y, Chu J, Li C. PIF4-mediated activation of YUCCA8 expression integrates temperature
422
into the auxin pathway in regulating arabidopsis hypocotyl growth. PLoS Genet. 2012. 8 (3): e1002594.
423
39. Spitschan M, Aguirre GK, Brainard DH, Sweeney AM. Variation of outdoor illumination as a function of
424
solar elevation and light pollution. Sci Rep. 2016. 6: 26756.
425
40. Viczian A, Kircher S, Fejes E, Millar AJ, Schafer E, Kozma-Bognar L, Nagy F. Functional characterization
426
of phytochrome interacting factor 3 for the Arabidopsis thaliana circadian clockwork. Plant and Cell
427
Physiology. 2005. 46: 1591–1602.
428
41. Yakir E, Hilman D, Harir Y, Green RM. Regulation of output from the plant circadian clock. FEBS J. 2007.
429
274 (2): 335-45.
430
42. Zhu L, Bu Q, Xu X, Paik I, Huang X, Hoecker U, Deng XW, Huq E. CUL4 forms an E3 ligase with COP1
431
and SPA to promote light-induced degradation of PIF1. Nature Communications. 2015. 6: 7245.
432
43. Zhu JY, Oh E, Wang T, Wang ZY. TOC1-PIF4 interaction mediates the circadian gating of thermoresponsive
433
growth in Arabidopsis. Nat Commun. 2016. 7:13692.
434 435
44.© 2019 by the authors. Submitted for possible open access publication under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).